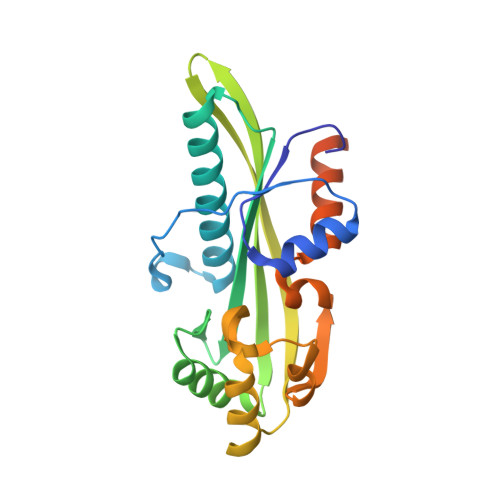
Biochemical and structural studies of conserved maf proteins revealed nucleotide pyrophosphatases with a preference for modified nucleotides.
Tchigvintsev, A., Tchigvintsev, D., Flick, R., Popovic, A., Dong, A., Xu, X., Brown, G., Lu, W., Wu, H., Cui, H., Dombrowski, L., Joo, J.C., Beloglazova, N., Min, J., Savchenko, A., Caudy, A.A., Rabinowitz, J.D., Murzin, A.G., Yakunin, A.F.(2013) Chem Biol 20: 1386-1398
- PubMed: 24210219
- DOI: https://doi.org/10.1016/j.chembiol.2013.09.011
- Primary Citation Related Structures:
2P5X, 4HEB, 4JHC, 4LU1 - PubMed Abstract:
Maf (for multicopy associated filamentation) proteins represent a large family of conserved proteins implicated in cell division arrest but whose biochemical activity remains unknown. Here, we show that the prokaryotic and eukaryotic Maf proteins exhibit nucleotide pyrophosphatase activity against 5-methyl-UTP, pseudo-UTP, 5-methyl-CTP, and 7-methyl-GTP, which represent the most abundant modified bases in all organisms, as well as against canonical nucleotides dTTP, UTP, and CTP. Overexpression of the Maf protein YhdE in E. coli cells increased intracellular levels of dTMP and UMP, confirming that dTTP and UTP are the in vivo substrates of this protein. Crystal structures and site-directed mutagenesis of Maf proteins revealed the determinants of their activity and substrate specificity. Thus, pyrophosphatase activity of Maf proteins toward canonical and modified nucleotides might provide the molecular mechanism for a dual role of these proteins in cell division arrest and house cleaning.
- Department of Chemical Engineering and Applied Chemistry, University of Toronto, Toronto, ON M5S 3E5, Canada.
Organizational Affiliation: