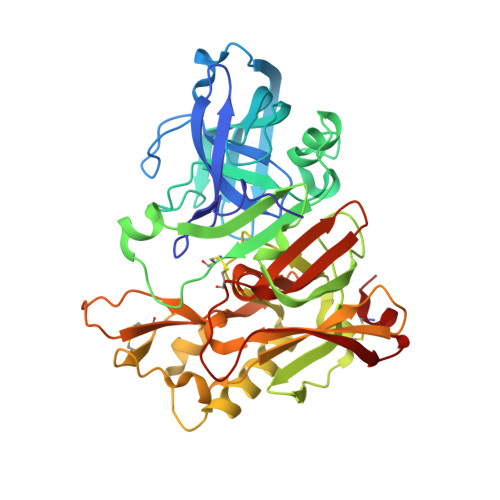
Design, synthesis, and X-ray structure of potent memapsin 2 (beta-secretase) inhibitors with isophthalamide derivatives as the P2-P3-ligands.
Ghosh, A.K., Kumaragurubaran, N., Hong, L., Kulkarni, S.S., Xu, X., Chang, W., Weerasena, V., Turner, R., Koelsch, G., Bilcer, G., Tang, J.(2007) J Med Chem 50: 2399-2407
- PubMed: 17432843 Search on PubMed
- DOI: https://doi.org/10.1021/jm061338s
- Primary Citation Related Structures:
2P4J - PubMed Abstract:
Structure-based design and synthesis of a number of potent and selective memapsin 2 inhibitors are described. These inhibitors were designed based upon the X-ray structure of memapsin 2-bound inhibitor 3 that incorporates methylsulfonyl alanine as the P2-ligand and a substituted pyrazole as the P3-ligand. Of particular importance, we examined the ability of the substituted isophthalic acid amide derivative to mimic the key interactions in the S2-S3 regions of the enzyme active sites of 3-bound memapsin 2. We investigated various substituted phenylethyl, alpha-methylbenzyl, and oxazolylmethyl groups as the P3-ligands. A number of inhibitors exhibited very potent inhibitory activity against mempasin 2 and good selectivity against memapsin 1. Inhibitor 5d has shown low nanomolar enzyme inhibitory potency (Ki=1.1 nM) and very good cellular inhibitory activity (IC50=39 nM). Furthermore, in a preliminary study, inhibitor 5d has shown 30% reduction of Abeta40 production in transgenic mice after a single intraperitoneal administration (8 mg/kg). A protein-ligand X-ray crystal structure of 5d-bound memapsin 2 provided vital molecular insight that can serve as an important guide to further design of novel inhibitors.
- Department of Chemistry, Purdue University, West Lafayette, Indiana 47907, USA. akghosh@purdue.edu
Organizational Affiliation: