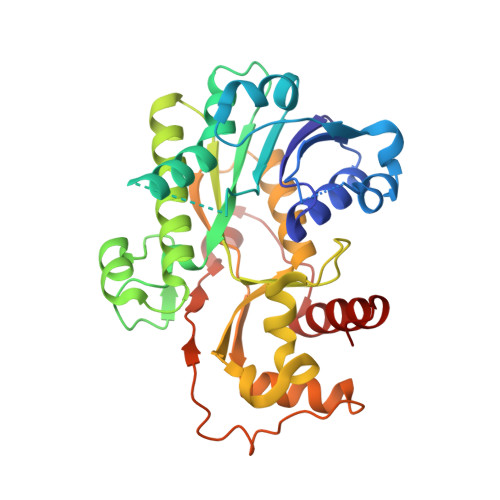
Crystal Structure of Vestitone Reductase from Alfalfa (Medicago sativa L.).
Shao, H., Dixon, R.A., Wang, X.(2007) J Mol Biology 369: 265-276
- PubMed: 17433362 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.03.040
- Primary Citation Related Structures:
2P4H - PubMed Abstract:
Isoflavonoids are commonly found in leguminous plants, where they play important roles in plant defense and have significant health benefits for animals and humans. Vestitone reductase catalyzes a stereospecific NADPH-dependent reduction of (3R)-vestitone in the biosynthesis of the antimicrobial isoflavonoid phytoalexin medicarpin. The crystal structure of alfalfa (Medicago sativa L.) vestitone reductase has been determined at 1.4 A resolution. The structure contains a classic Rossmann fold domain in the N terminus and a small C-terminal domain. Sequence and structural analysis showed that vestitone reductase is a member of the short-chain dehydrogenase/reductase (SDR) superfamily despite the low levels of sequence identity, and the prominent structural differences from other SDR enzymes with known structures. The putative binding sites for the co-factor NADPH and the substrate (3R)-vestitone were defined and located in a large cleft formed between the N and C-terminal domains of enzyme. Potential key residues for enzyme activity were also identified, including the catalytic triad Ser129-Tyr164-Lys168. A molecular docking study showed that (3R)-vestitone, but not the (3S) isomer, forms favored interactions with the co-factor and catalytic triad, thus providing an explanation for the enzyme's strict substrate stereo-specificity.
- Plant Biology Division, Samuel Roberts Noble Foundation, 2510 Sam Noble Parkway, Ardmore, OK 73401, USA.
Organizational Affiliation: