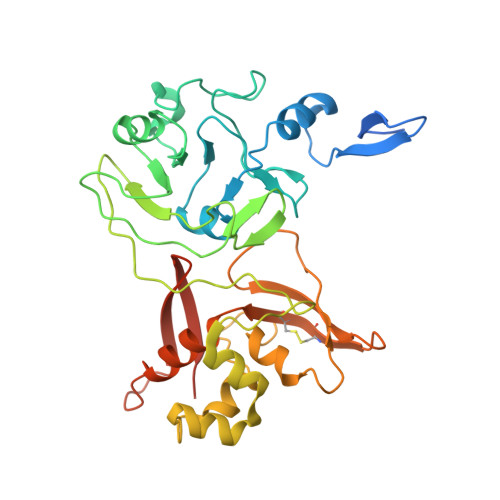
Crystal structure of a monomeric form of severe acute respiratory syndrome coronavirus endonuclease nsp15 suggests a role for hexamerization as an allosteric switch.
Joseph, J.S., Saikatendu, K.S., Subramanian, V., Neuman, B.W., Buchmeier, M.J., Stevens, R.C., Kuhn, P.(2007) J Virol 81: 6700-6708
- PubMed: 17409150 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.02817-06
- Primary Citation Related Structures:
2OZK - PubMed Abstract:
Mature nonstructural protein-15 (nsp15) from the severe acute respiratory syndrome coronavirus (SARS-CoV) contains a novel uridylate-specific Mn2+-dependent endoribonuclease (NendoU). Structure studies of the full-length form of the obligate hexameric enzyme from two CoVs, SARS-CoV and murine hepatitis virus, and its monomeric homologue, XendoU from Xenopus laevis, combined with mutagenesis studies have implicated several residues in enzymatic activity and the N-terminal domain as the major determinant of hexamerization. However, the tight link between hexamerization and enzyme activity in NendoUs has remained an enigma. Here, we report the structure of a trimmed, monomeric form of SARS-CoV nsp15 (residues 28 to 335) determined to a resolution of 2.9 A. The catalytic loop (residues 234 to 249) with its two reactive histidines (His 234 and His 249) is dramatically flipped by approximately 120 degrees into the active site cleft. Furthermore, the catalytic nucleophile Lys 289 points in a diametrically opposite direction, a consequence of an outward displacement of the supporting loop (residues 276 to 295). In the full-length hexameric forms, these two loops are packed against each other and are stabilized by intimate intersubunit interactions. Our results support the hypothesis that absence of an adjacent monomer due to deletion of the hexamerization domain is the most likely cause for disruption of the active site, offering a structural basis for why only the hexameric form of this enzyme is active.
- Department of Cell Biology, 10550 N. Torrey Pines Road, CB265, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: