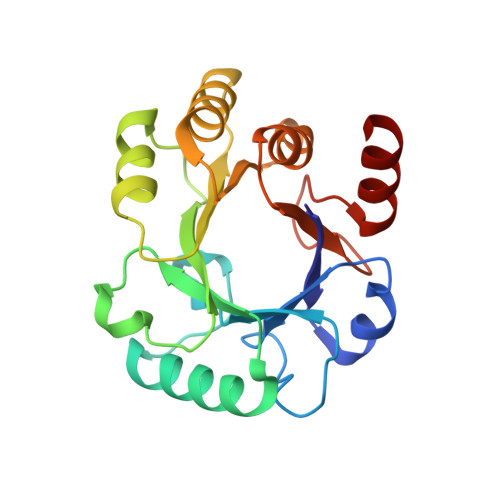
Structure and lability of archaeal dehydroquinase.
Smith, N.N., Gallagher, D.T.(2008) Acta Crystallogr Sect F Struct Biol Cryst Commun 64: 886-892
- PubMed: 18931429 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309108028546
- Primary Citation Related Structures:
2OX1 - PubMed Abstract:
Multiple sequence alignments of type I 3-dehydroquinate dehydratases (DQs; EC 4.2.1.10) show that archaeal DQs have shorter helical regions than bacterial orthologs of known structure. To investigate this feature and its relation to thermostability, the structure of the Archaeoglobus fulgidus (Af) DQ dimer was determined at 2.33 A resolution and its denaturation temperature was measured in vitro by circular dichroism (CD) and differential scanning calorimetry (DSC). This structure, a P2(1)2(1)2(1) crystal form with two 45 kDa dimers in the asymmetric unit, is the first structural representative of an archaeal DQ. Denaturation occurs at 343 +/- 3 K at both low and high ionic strength and at 349 K in the presence of the substrate analog tartrate. Since the growth optimum of the organism is 356 K, this implies that the protein maintains its folded state through the participation of additional factors in vivo. The (betaalpha)(8) fold is compared with those of two previously determined type I DQ structures, both bacterial (Salmonella and Staphylococcus), which had sequence identities of approximately 30% with AfDQ. Although the overall folds are the same, there are many differences in secondary structure and ionic features; the archaeal protein has over twice as many salt links per residue. The archaeal DQ is smaller than its bacterial counterparts and lower in regular secondary structure, with its eight helices being an average of one turn shorter. In particular, two of the eight normally helical regions (the exterior of the barrel) are mostly nonhelical in AfDQ, each having only a single turn of 3(10)-helix flanked by beta-strand and coil. These ;protohelices' are unique among evolutionarily close members of the (betaalpha)(8)-fold superfamily. Structural features that may contribute to stability, in particular ionic factors, are examined and the implications of having a T(m) below the organism's growth temperature are considered.
- National Institute of Standards and Technology, Biochemical Science Division, Gaithersburg, MD 20899-8312, USA.
Organizational Affiliation: