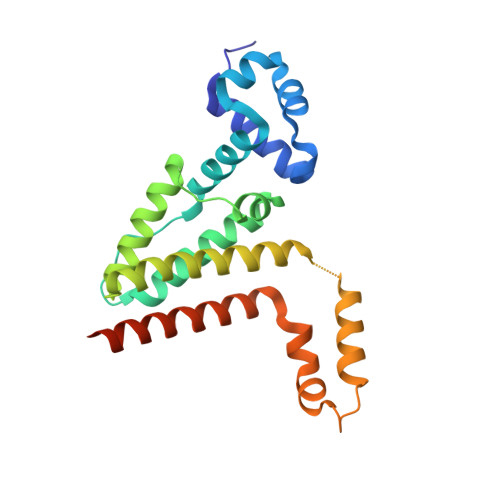
Crystal structures of the Streptomyces coelicolor TetR-like protein ActR alone and in complex with actinorhodin or the actinorhodin biosynthetic precursor (S)-DNPA.
Willems, A.R., Tahlan, K., Taguchi, T., Zhang, K., Lee, Z.Z., Ichinose, K., Junop, M.S., Nodwell, J.R.(2008) J Mol Biology 376: 1377-1387
- PubMed: 18207163 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.12.061
- Primary Citation Related Structures:
2OPT, 3B6A, 3B6C - PubMed Abstract:
Actinorhodin, an antibiotic produced by Streptomyces coelicolor, is exported from the cell by the ActA efflux pump. actA is divergently transcribed from actR, which encodes a TetR-like transcriptional repressor. We showed previously that ActR represses transcription by binding to an operator from the actA/actR intergenic region. Importantly, actinorhodin itself or various actinorhodin biosynthetic intermediates can cause ActR to dissociate from its operator, leading to derepression. This suggests that ActR may mediate timely self-resistance to an endogenously produced antibiotic by responding to one of its biosynthetic precursors. Here, we report the structural basis for this precursor-mediated derepression with crystal structures of homodimeric ActR by itself and in complex with either actinorhodin or the actinorhodin biosynthetic intermediate (S)-DNPA [4-dihydro-9-hydroxy-1-methyl-10-oxo-3-H-naphtho-[2,3-c]-pyran-3-(S)-acetic acid]. The ligand-binding tunnel in each ActR monomer has a striking hydrophilic/hydrophobic/hydrophilic arrangement of surface residues that accommodate either one hexacyclic actinorhodin molecule or two back-to-back tricyclic (S)-DNPA molecules. Moreover, our work also reveals the strongest structural evidence to date that TetR-mediated antibiotic resistance may have been acquired from an antibiotic-producer organism.
- Department of Biochemistry and Biomedical Sciences, McMaster University, 1200 Main Street West, Hamilton, Ontario, Canada.
Organizational Affiliation: