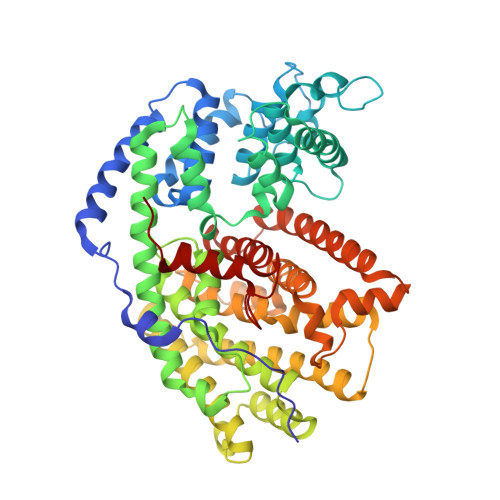
Structure of limonene synthase, a simple model for terpenoid cyclase catalysis.
Hyatt, D.C., Youn, B., Zhao, Y., Santhamma, B., Coates, R.M., Croteau, R.B., Kang, C.(2007) Proc Natl Acad Sci U S A 104: 5360-5365
- PubMed: 17372193 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0700915104
- Primary Citation Related Structures:
2ONG, 2ONH - PubMed Abstract:
The crystal structure of (4S)-limonene synthase from Mentha spic ata, a metal ion-dependent monoterpene cyclase that catalyzes the coupled isomerization and cyclization of geranyl diphosphate, is reported at 2.7-A; resolution in two forms liganded to the substrate and intermediate analogs, 2-fluorogeranyl diphosphate and 2-fluorolinalyl diphosphate, respectively. The implications of these findings are described for domain interactions in the homodimer and for changes in diphosphate-metal ion coordination and substrate binding conformation in the course of the multistep reaction.
- Institute of Biological Chemistry, Washingston State University, Pullman, WA 99164-6340, USA.
Organizational Affiliation: