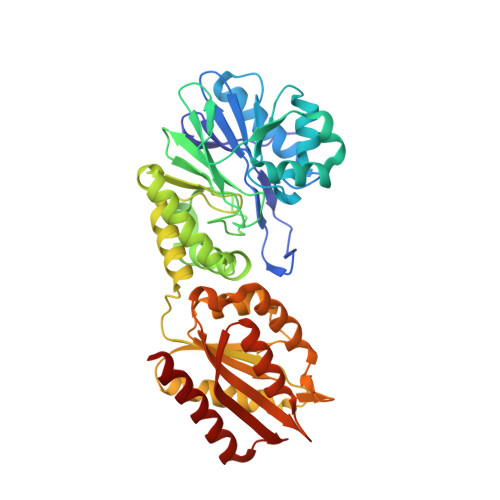
Structure of coenzyme F420H2 oxidase (FprA), a di-iron flavoprotein from methanogenic Archaea catalyzing the reduction of O2 to H2O.
Seedorf, H., Hagemeier, C.H., Shima, S., Thauer, R.K., Warkentin, E., Ermler, U.(2007) FEBS J 274: 1588-1599
- PubMed: 17480207
- DOI: https://doi.org/10.1111/j.1742-4658.2007.05706.x
- Primary Citation Related Structures:
2OHH, 2OHI, 2OHJ - PubMed Abstract:
The di-iron flavoprotein F(420)H(2) oxidase found in methanogenic Archaea catalyzes the four-electron reduction of O(2) to 2H(2)O with 2 mol of reduced coenzyme F(420)(7,8-dimethyl-8-hydroxy-5-deazariboflavin). We report here on crystal structures of the homotetrameric F(420)H(2) oxidase from Methanothermobacter marburgensis at resolutions of 2.25 A, 2.25 A and 1.7 A, respectively, from which an active reduced state, an inactive oxidized state and an active oxidized state could be extracted. As found in structurally related A-type flavoproteins, the active site is formed at the dimer interface, where the di-iron center of one monomer is juxtaposed to FMN of the other. In the active reduced state [Fe(II)Fe(II)FMNH(2)], the two irons are surrounded by four histidines, one aspartate, one glutamate and one bridging aspartate. The so-called switch loop is in a closed conformation, thus preventing F(420) binding. In the inactive oxidized state [Fe(III)FMN], the iron nearest to FMN has moved to two remote binding sites, and the switch loop is changed to an open conformation. In the active oxidized state [Fe(III)Fe(III)FMN], both irons are positioned as in the reduced state but the switch loop is found in the open conformation as in the inactive oxidized state. It is proposed that the redox-dependent conformational change of the switch loop ensures alternate complete four-electron O(2) reduction and redox center re-reduction. On the basis of the known Si-Si stereospecific hydride transfer, F(420)H(2) was modeled into the solvent-accessible pocket in front of FMN. The inactive oxidized state might provide the molecular basis for enzyme inactivation by long-term O(2) exposure observed in some members of the FprA family.
- Max Planck Institute for Terrestrial Microbiology, Marburg, Germany.
Organizational Affiliation: