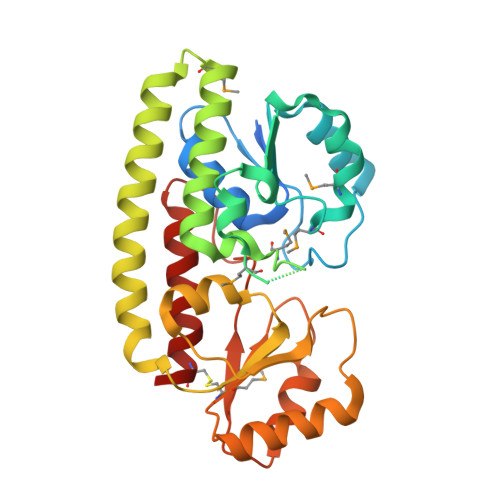
Structural Analysis of ABC-family Periplasmic Zinc Binding Protein Provides New Insights Into Mechanism of Ligand Uptake and Release
Chandra, B.R., Yogavel, M., Sharma, A.(2007) J Mol Biology 367: 970-982
- PubMed: 17306297 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2007.01.041
- Primary Citation Related Structures:
2OGW - PubMed Abstract:
ATP-binding cassette superfamily of periplasmic metal transporters are known to be vital for maintaining ion homeostasis in several pathogenic and non-pathogenic bacteria. We have determined crystal structure of the high-affinity zinc transporter ZnuA from Escherichia coli at 1.8 A resolution. This structure represents the first native (non-recombinant) protein structure of a periplasmic metal binding protein. ZnuA reveals numerous conformational features, which occur either in Zn(2+) or in Mn(2+) transporters, and presents a unique conformational state. A comprehensive comparison of ZnuA with other periplasmic ligand binding protein structures suggests vital mechanistic differences between bound and release states of metal transporters. The key new attributes in ZnuA include a C-domain disulfide bond, an extra alpha-helix proximal to the highly charged metal chelating mobile loop region, alternate conformations of secondary shell stabilizing residues at the metal binding site, and domain movements potentially controlled by salt bridges. Based on in-depth structural analyses of five metal binding transporters, we present here a mechanistic model termed as "partial domain slippage" for binding and release of Zn(2+).
- Structural and Computational Biology Group, International Centre for Genetic Engineering and Biotechnology, New Delhi, India.
Organizational Affiliation: