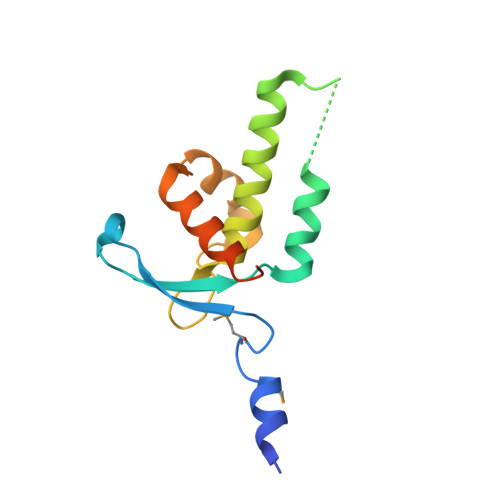
A conserved spiral structure for highly diverged phage tail assembly chaperones.
Pell, L.G., Cumby, N., Clark, T.E., Tuite, A., Battaile, K.P., Edwards, A.M., Chirgadze, N.Y., Davidson, A.R., Maxwell, K.L.(2013) J Mol Biology 425: 2436-2449
- PubMed: 23542344 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2013.03.035
- Primary Citation Related Structures:
2OB9 - PubMed Abstract:
Tail assembly chaperones (TACs) are a family of proteins likely required for the morphogenesis of all long-tailed phages. In this study, we determined the crystal structure of gp13, the TAC of phage HK97. This structure is similar to that of the TAC from the Lactococcus phage p2 and two unannotated structures of likely TACs encoded in prophage-derived regions of Bacillus subtilis and Bacillus stearothermophilus. Despite the high sequence divergence of these proteins, gp13 forms a ring structure with similar dimensions to the spirals observed in the crystal lattices of these other proteins. Remarkably, these similar quaternary structures are formed through very different interprotomer interactions. We present functional data supporting the biological relevance of these spiral structures and propose that spiral formation has been the primary requirement for these proteins during evolution. This study presents an unusual example of diverged protein sequences and oligomerization mechanisms in the presence of conserved quaternary structure.
- Department of Molecular Genetics and Donnelly Centre for Cellular and Biomolecular Research, University of Toronto, Toronto, ON, Canada M5S 3E1.
Organizational Affiliation: