Crystal structure of calmodulin-neuronal nitric oxide synthase complex
Valentine, K.G., Ng, H.L., Schneeweis, L., Kranz, J.K., Frederick, K.K., Alber, T., Wand, A.J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Currently 2O60 does not have a validation slider image.
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Calmodulin | 148 | Gallus gallus | Mutation(s): 0 Gene Names: CALM | 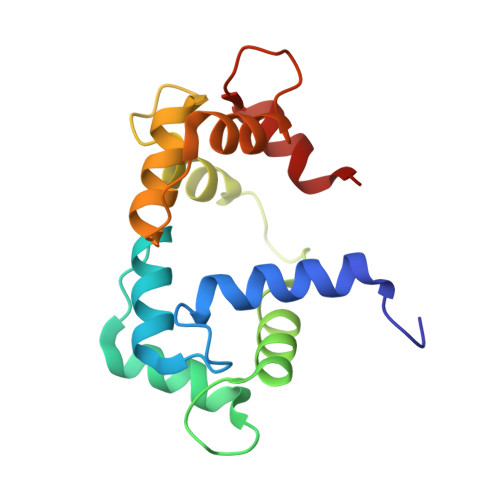 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P62149 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Peptide corresponding to calmodulin binding domain of neuronal nitric oxide synthase | 24 | N/A | Mutation(s): 0 EC: 1.14.13.39 | 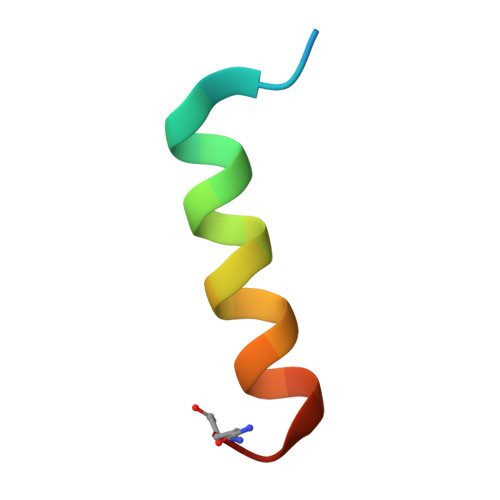 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9Z0J4 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CA Download:Ideal Coordinates CCD File | C [auth A], D [auth A], E [auth A], F [auth A] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 67.126 | α = 90 |
| b = 32.93 | β = 93.3 |
| c = 73.83 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| Blu-Ice | data collection |
| MOSFLM | data reduction |
| SCALA | data scaling |
| EPMR | phasing |
Currently 2O60 does not have a validation slider image.