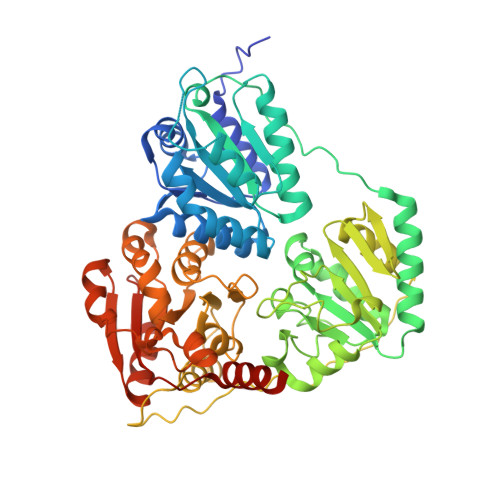
The crystal structure of phenylpyruvate decarboxylase from Azospirillum brasilense at 1.5 A resolution. Implications for its catalytic and regulatory mechanism.
Versees, W., Spaepen, S., Vanderleyden, J., Steyaert, J.(2007) FEBS J 274: 2363-2375
- PubMed: 17403037 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2007.05771.x
- Primary Citation Related Structures:
2NXW - PubMed Abstract:
Phenylpyruvate decarboxylase (PPDC) of Azospirillum brasilense, involved in the biosynthesis of the plant hormone indole-3-acetic acid and the antimicrobial compound phenylacetic acid, is a thiamine diphosphate-dependent enzyme that catalyses the nonoxidative decarboxylation of indole- and phenylpyruvate. Analogous to yeast pyruvate decarboxylases, PPDC is subject to allosteric substrate activation, showing sigmoidal v versus [S] plots. The present paper reports the crystal structure of this enzyme determined at 1.5 A resolution. The subunit architecture of PPDC is characteristic for other members of the pyruvate oxidase family, with each subunit consisting of three domains with an open alpha/beta topology. An active site loop, bearing the catalytic residues His112 and His113, could not be modelled due to flexibility. The biological tetramer is best described as an asymmetric dimer of dimers. A cysteine residue that has been suggested as the site for regulatory substrate binding in yeast pyruvate decarboxylase is not conserved, requiring a different mechanism for allosteric substrate activation in PPDC. Only minor changes occur in the interactions with the cofactors, thiamine diphosphate and Mg2+, compared to pyruvate decarboxylase. A greater diversity is observed in the substrate binding pocket accounting for the difference in substrate specificity. Moreover, a catalytically important glutamate residue conserved in nearly all decarboxylases is replaced by a leucine in PPDC. The consequences of these differences in terms of the catalytic and regulatory mechanism of PPDC are discussed.
- Department of Ultrastructure, Vrije Universiteit Brussel, Brussels, Belgium. wversees@vub.ac.be
Organizational Affiliation: