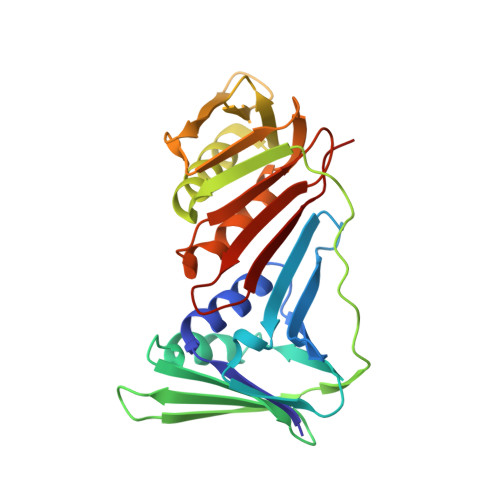
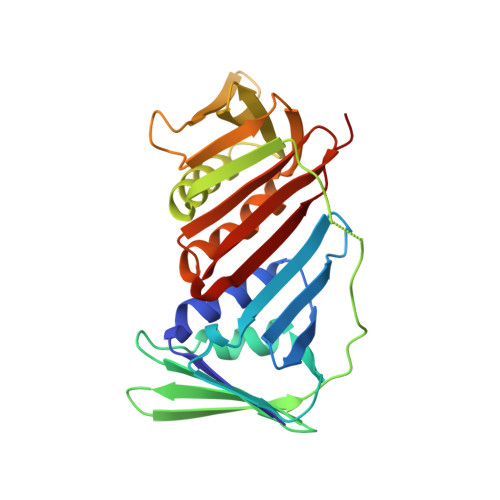
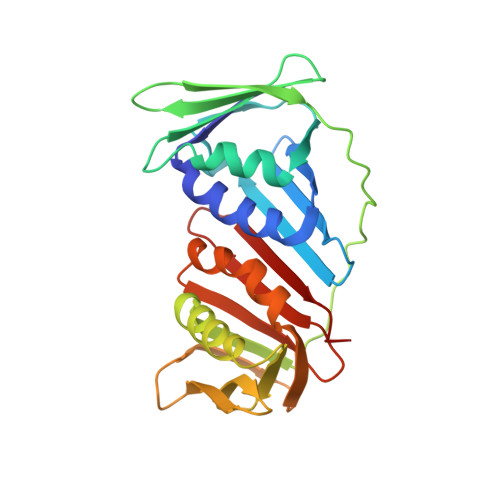
Structures of monomeric, dimeric and trimeric PCNA: PCNA-ring assembly and opening.
Hlinkova, V., Xing, G., Bauer, J., Shin, Y.J., Dionne, I., Rajashankar, K.R., Bell, S.D., Ling, H.(2008) Acta Crystallogr D Biol Crystallogr 64: 941-949
- PubMed: 18703842
- DOI: https://doi.org/10.1107/S0907444908021665
- Primary Citation Related Structures:
2IJX, 2IO4, 2NTI - PubMed Abstract:
DNA sliding clamps form an oligomeric ring encircling DNA and serve as a moving platform for DNA-processing proteins. The opening and closing of a sliding-clamp ring is essential to load the clamp onto DNA in order to perform its functions. The molecular details of how clamp rings open and enclose DNA are still not clear. Three PCNA homologues have been found in Sulfolobus solfataricus which form a heterotrimer. Taking advantage of their hetero-oligomeric nature, the structures of the PCNAs in monomeric PCNA3, dimeric PCNA1-PCNA2 and trimeric PCNA1-PCNA2-PCNA3 forms were determined at resolutions of 2.6-1.9 A. The distinct oligomeric structures represent different stages in ring formation, which were verified in solution by ultracentrifugation analysis. The heterodimer opens in a V-shape of 130 degrees , while the heterotrimers form a ring with a 120 degrees rotation between monomers. The association of a rigid PCNA3 monomer with an opened PCNA1-PCNA2 heterodimer closes the ring and introduces a spring tension in the PCNA1-PCNA2 interface, thus bending the nine-stranded intermolecular beta-sheet to fit the 120 degrees rotation. The release of the spring tension as PCNA3 dissociates from the ring may facilitate ring opening. The structural features in different assemblies present a molecular model for clamp ring assembly and opening.
- Department of Biochemistry, University of Western Ontario, London, Ontario N6A 5C1, Canada.
Organizational Affiliation: