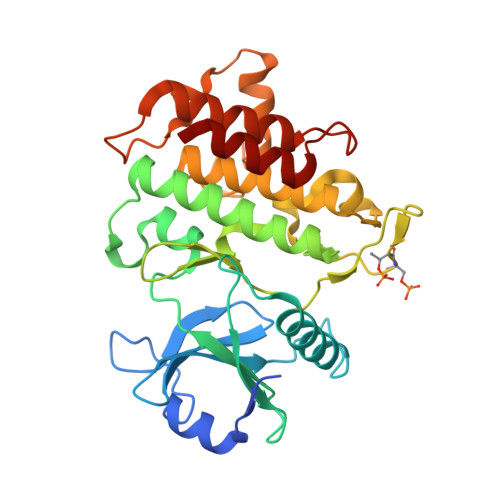
Crystal structures of IRAK-4 kinase in complex with inhibitors: a serine/threonine kinase with tyrosine as a gatekeeper.
Wang, Z., Liu, J., Sudom, A., Ayres, M., Li, S., Wesche, H., Powers, J.P., Walker, N.P.C.(2006) Structure 14: 1835-1844
- PubMed: 17161373 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2006.11.001
- Primary Citation Related Structures:
2NRU, 2NRY - PubMed Abstract:
Interleukin-1 (IL-1) receptor-associated kinase-4 (IRAK-4) is a serine/threonine kinase that plays an essential role in signal transduction by Toll/IL-1 receptors (TIRs). Here, we report the crystal structures of the phosphorylated human IRAK-4 kinase domain in complex with a potent inhibitor and with staurosporine to 2.0 and 2.2 A, respectively. The structures reveal that IRAK-4 has a unique tyrosine gatekeeper residue that interacts with the conserved glutamate from helix alphaC. Consequently, helix alphaC is "pulled in" to maintain the active orientation, and the usual pre-existing hydrophobic back pocket of the ATP-binding site is abolished. The peptide substrate-binding site is more open when compared with other protein kinases due to a marked movement of helix alphaG. The pattern of phosphate ligand interactions in the activation loop bears a close resemblance to that of a tyrosine kinase. Our results provide insights into IRAK-4 function and the design of selective inhibitors.
- Department of Molecular Structure, Amgen, Inc., 1120 Veterans Boulevard, South San Francisco, California 94080, USA. zwang@amgen.com <zwang@amgen.com>
Organizational Affiliation: