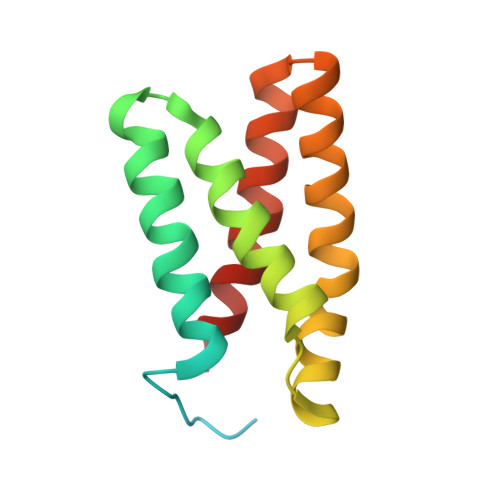
Structural characterization of the interaction of mTOR with phosphatidic acid and a novel class of inhibitor: compelling evidence for a central role of the FRB domain in small molecule-mediated regulation of mTOR.
Veverka, V., Crabbe, T., Bird, I., Lennie, G., Muskett, F.W., Taylor, R.J., Carr, M.D.(2008) Oncogene 27: 585-595
- PubMed: 17684489 Search on PubMed
- DOI: https://doi.org/10.1038/sj.onc.1210693
- Primary Citation Related Structures:
2NPU - PubMed Abstract:
The mammalian target of rapamycin (mTOR) is a large, multidomain protein kinase, which plays a central role in the regulation of cell growth and has recently emerged as an essential target of survival signals in many types of human cancer cells. Here, we report the solution structures of complexes formed between the FKBP12-rapamycin binding (FRB) domain of mTOR and phosphatidic acid, an important cellular activator of the kinase, and between the FRB domain and a novel inhibitor (HTS-1). The overall structure of the FRB domain is very similar to that seen in the ternary complex formed with FKBP12 and the immunosuppressive drug rapamycin; however, there are significant changes within the rapamycin-binding site with important consequences for rational drug design. The surface of the FRB domain contains a number of distinctive features that have previously escaped attention, including a potential new regulatory site on the opposite face to that involved in the binding of rapamycin, which displays the features expected for a specific binding site for a small molecule. The interaction sites for phosphatidic acid and HTS-1 were found to closely match the site responsible for rapamycin binding. In addition, the structures determined for the FRB-phosphatidic acid and FRB-HTS-1 complexes revealed a striking similarity between the conformations of buried portions of the ligands and that seen for the rapamycin backbone in contact with the domain. Our findings further highlight the importance of the FRB domain in small molecule-mediated regulation of mTOR, demonstrate the ability to identify novel inhibitors of mTOR that bind tightly to the rapamycin-binding site in the absence of FKBP12, and identify a potential new regulatory site that may be exploited in the design of new anticancer drugs.
- Department of Biochemistry, University of Leicester, Leicester, UK.
Organizational Affiliation: