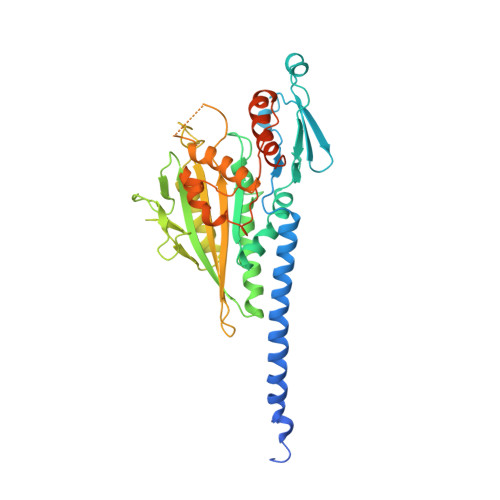
Direction determination in the minus-end-directed kinesin motor ncd.
Sablin, E.P., Case, R.B., Dai, S.C., Hart, C.L., Ruby, A., Vale, R.D., Fletterick, R.J.(1998) Nature 395: 813-816
- PubMed: 9796817
- DOI: https://doi.org/10.1038/27463
- Primary Citation of Related Structures:
2NCD - PubMed Abstract:
Motor proteins of the kinesin superfamily transport intracellular cargo along microtubules. Although different kinesin proteins share 30-50% amino-acid identity in their motor catalytic cores, some move to the plus end of microtubules whereas others travel in the opposite direction. Crystal structures of the catalytic cores of conventional kinesin (a plus-end-directed motor involved in organelle transport) and ncd (a minus-end-directed motor involved in chromosome segregation) are nearly identical; therefore, the structural basis for their opposite directions of movement is unknown. Here we show that the ncd 'neck' made up of 13 class-specific residues next to the superfamily-conserved catalytic core, is essential for minus-end-directed motility, as mutagenesis of these neck residues reverses the direction of ncd motion. By solving the 2.5 A structure of a functional ncd dimer, we show that the ncd neck (a coiled-coil) differs from the corresponding region in the kinesin neck (an interrupted beta-strand), although both necks interact with similar elements in the catalytic cores. The distinct neck architectures also confer different symmetries to the ncd and kinesin dimers and position these motors with appropriate directional bias on the microtubule.
- Department of Biochemistry/Biophysics, University of California, San Francisco 94143, USA.
Organizational Affiliation: