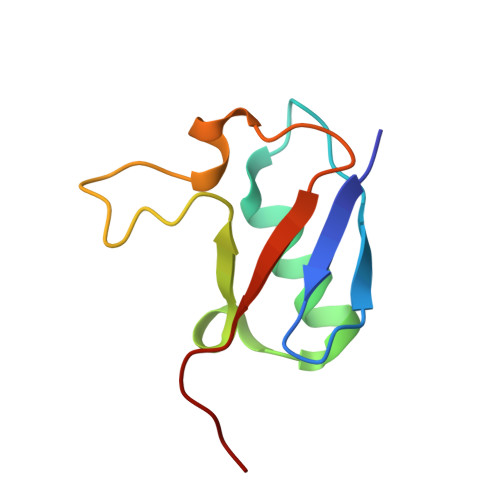
Observing a late folding intermediate of Ubiquitin at atomic resolution by NMR
Surana, P., Das, R.(2016) Protein Sci 25: 1438-1450
- PubMed: 27111887 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2940
- Primary Citation Related Structures:
2NBD, 2NBE - PubMed Abstract:
The study of intermediates in the protein folding pathway provides a wealth of information about the energy landscape. The intermediates also frequently initiate pathogenic fibril formations. While observing the intermediates is difficult due to their transient nature, extreme conditions can partially unfold the proteins and provide a glimpse of the intermediate states. Here, we observe the high resolution structure of a hydrophobic core mutant of Ubiquitin at an extreme acidic pH by nuclear magnetic resonance (NMR) spectroscopy. In the structure, the native secondary and tertiary structure is conserved for a major part of the protein. However, a long loop between the beta strands β3 and β5 is partially unfolded. The altered structure is supported by fluorescence data and the difference in free energies between the native state and the intermediate is reflected in the denaturant induced melting curves. The unfolded region includes amino acids that are critical for interaction with cofactors as well as for assembly of poly-Ubiquitin chains. The structure at acidic pH resembles a late folding intermediate of Ubiquitin and indicates that upon stabilization of the protein's core, the long loop converges on the core in the final step of the folding process.
- National Centre for Biological Sciences, Tata Institute of Fundamental Research, Bengaluru, 560065, Karnataka, India.
Organizational Affiliation: