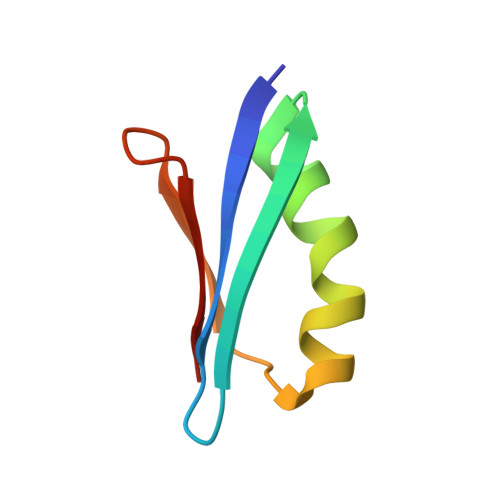
Improved in-cell structure determination of proteins at near-physiological concentration
Ikeya, T., Hanashima, T., Hosoya, S., Shimazaki, M., Ikeda, S., Mishima, M., Guentert, P., Ito, Y.(2016) Sci Rep 6: 38312-38312
- PubMed: 27910948 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep38312
- Primary Citation Related Structures:
2N9K, 2N9L - PubMed Abstract:
Investigating three-dimensional (3D) structures of proteins in living cells by in-cell nuclear magnetic resonance (NMR) spectroscopy opens an avenue towards understanding the structural basis of their functions and physical properties under physiological conditions inside cells. In-cell NMR provides data at atomic resolution non-invasively, and has been used to detect protein-protein interactions, thermodynamics of protein stability, the behavior of intrinsically disordered proteins, etc. in cells. However, so far only a single de novo 3D protein structure could be determined based on data derived only from in-cell NMR. Here we introduce methods that enable in-cell NMR protein structure determination for a larger number of proteins at concentrations that approach physiological ones. The new methods comprise (1) advances in the processing of non-uniformly sampled NMR data, which reduces the measurement time for the intrinsically short-lived in-cell NMR samples, (2) automatic chemical shift assignment for obtaining an optimal resonance assignment, and (3) structure refinement with Bayesian inference, which makes it possible to calculate accurate 3D protein structures from sparse data sets of conformational restraints. As an example application we determined the structure of the B1 domain of protein G at about 250 μM concentration in living E. coli cells.
- Department of Chemistry, Graduate School of Science and Engineering, Tokyo Metropolitan University, Tokyo, 192-0397, Japan.
Organizational Affiliation: