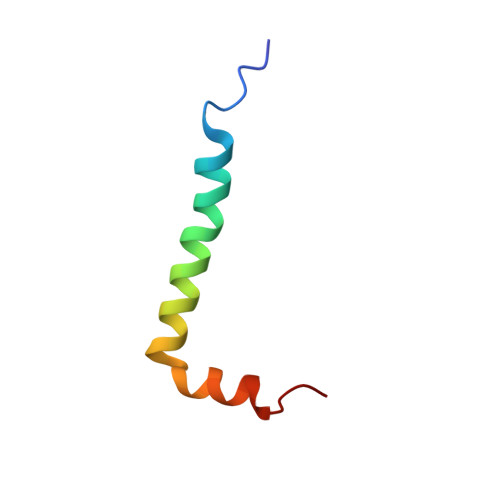
Structure and Mechanism of the Influenza A M218-60 Dimer of Dimers.
Andreas, L.B., Reese, M., Eddy, M.T., Gelev, V., Ni, Q.Z., Miller, E.A., Emsley, L., Pintacuda, G., Chou, J.J., Griffin, R.G.(2015) J Am Chem Soc 137: 14877-14886
- PubMed: 26218479 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.5b04802
- Primary Citation Related Structures:
2N70 - PubMed Abstract:
We report a magic angle spinning (MAS) NMR structure of the drug-resistant S31N mutation of M218-60 from Influenza A. The protein was dispersed in diphytanoyl-sn-glycero-3-phosphocholine lipid bilayers, and the spectra and an extensive set of constraints indicate that M218-60 consists of a dimer of dimers. In particular, ∼280 structural constraints were obtained using dipole recoupling experiments that yielded well-resolved (13)C-(15)N, (13)C-(13)C, and (1)H-(15)N 2D, 3D, and 4D MAS spectra, all of which show cross-peak doubling. Interhelical distances were measured using mixed (15)N/(13)C labeling and with deuterated protein, MAS at ωr/2π = 60 kHz, ω0H/2π = 1000 MHz, and (1)H detection of methyl-methyl contacts. The experiments reveal a compact structure consisting of a tetramer composed of four transmembrane helices, in which two opposing helices are displaced and rotated in the direction of the membrane normal relative to a four-fold symmetric arrangement, yielding a two-fold symmetric structure. Side chain conformations of the important gating and pH-sensing residues W41 and H37 are found to differ markedly from four-fold symmetry. The rmsd of the structure is 0.7 Å for backbone heavy atoms and 1.1 Å for all heavy atoms. This two-fold symmetric structure is different from all of the previous structures of M2, many of which were determined in detergent and/or with shorter constructs that are not fully active. The structure has implications for the mechanism of H(+) transport since the distance between His and Trp residues on different helices is found to be short. The structure also exhibits two-fold symmetry in the vicinity of the binding site of adamantyl inhibitors, and steric constraints may explain the mechanism of the drug-resistant S31N mutation.
- Department of Chemistry and Francis Bitter Magnet Laboratory, Massachusetts Institute of Technology , 77 Massachusetts Avenue, Cambridge, Massachusetts 02139, United States.
Organizational Affiliation: