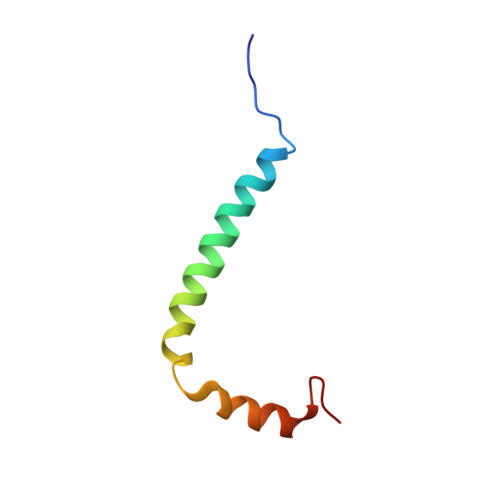
The Membrane Mimetic Affects the Spatial Structure and Mobility of EGFR Transmembrane and Juxtamembrane Domains.
Mineev, K.S., Panova, S.V., Bocharova, O.V., Bocharov, E.V., Arseniev, A.S.(2015) Biochemistry 54: 6295-6298
- PubMed: 26440883 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.5b00851
- Primary Citation Related Structures:
2N5S - PubMed Abstract:
The epidermal growth factor receptor (EGFR) is one of the most extensively studied receptor tyrosine kinases, as it is involved in a wide range of cellular processes and severe diseases. Recent works reveal that the single-helix transmembrane domains and cytoplasmic juxtamembrane regions play an important role in the receptor activation process. Here we present the results of our investigation of the spatial structure and mobility of the EGFR transmembrane domain and juxtamembrane regions in various membranelike environments, which shed light on the effects of the membrane physical properties and composition on the behavior of the juxtamembrane domain.
- Shemyakin-Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences RAS , str. Miklukho-Maklaya 16/10, Moscow, 117997 Russian Federation.
Organizational Affiliation: