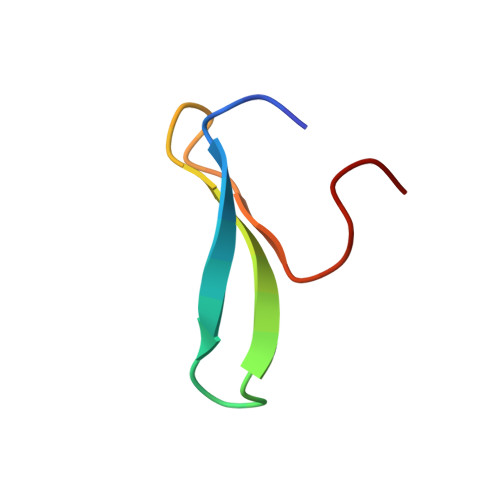
Structural Analysis of the Pin1-CPEB1 interaction and its potential role in CPEB1 degradation.
Schelhorn, C., Martin-Malpartida, P., Sunol, D., Macias, M.J.(2015) Sci Rep 5: 14990-14990
- PubMed: 26456073 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep14990
- Primary Citation Related Structures:
2N1O - PubMed Abstract:
The Cytoplasmic Polyadenylation Element Binding proteins are RNA binding proteins involved in the translational regulation of mRNA. During cell cycle progression, CPEB1 is labeled for degradation by phosphorylation-dependent ubiquitination by the SCF(β-TrCP) ligase. The peptidyl-prolyl isomerase Pin1 plays a key role in CPEB1 degradation. Conditioned by the cell cycle stage, CPEB1 and Pin1 interactions occur in a phosphorylation-independent or -dependent manner. CPEB1 contains six potential phosphorylatable Pin1 binding sites. Using a set of biophysical techniques, we discovered that the pS210 site is unique, since it displays binding activity not only to the WW domain but also to the prolyl-isomerase domain of Pin1. The NMR structure of the Pin1 WW-CPEB1 pS210 (PDB ID: 2n1o) reveals that the pSerPro motif is bound in trans configuration through contacts with amino acids located in the first turn of the WW domain and the conserved tryptophan in the β3-strand. NMR relaxation analyses of Pin1 suggest that inter-domain flexibility is conferred by the modulation of the interaction with peptides containing the pS210 site, which is essential for degradation.
- Institute for Research in Biomedicine (IRB Barcelona), The Barcelona Institute of Science and Technology (BIST), Baldiri Reixac 10, Barcelona, 08028, Spain.
Organizational Affiliation: