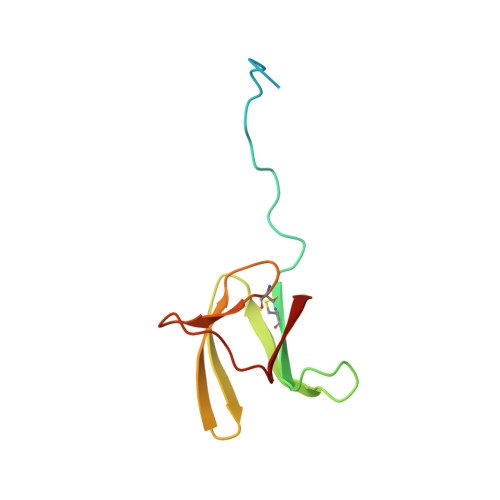
Structure Analysis Uncovers a Highly Diverse but Structurally Conserved Effector Family in Phytopathogenic Fungi.
de Guillen, K., Ortiz-Vallejo, D., Gracy, J., Fournier, E., Kroj, T., Padilla, A.(2015) PLoS Pathog 11: e1005228-e1005228
- PubMed: 26506000
- DOI: https://doi.org/10.1371/journal.ppat.1005228
- Primary Citation of Related Structures:
2MYV, 2MYW - PubMed Abstract:
Phytopathogenic ascomycete fungi possess huge effector repertoires that are dominated by hundreds of sequence-unrelated small secreted proteins. The molecular function of these effectors and the evolutionary mechanisms that generate this tremendous number of singleton genes are largely unknown. To get a deeper understanding of fungal effectors, we determined by NMR spectroscopy the 3-dimensional structures of the Magnaporthe oryzae effectors AVR1-CO39 and AVR-Pia. Despite a lack of sequence similarity, both proteins have very similar 6 β-sandwich structures that are stabilized in both cases by a disulfide bridge between 2 conserved cysteins located in similar positions of the proteins. Structural similarity searches revealed that AvrPiz-t, another effector from M. oryzae, and ToxB, an effector of the wheat tan spot pathogen Pyrenophora tritici-repentis have the same structures suggesting the existence of a family of sequence-unrelated but structurally conserved fungal effectors that we named MAX-effectors (Magnaporthe Avrs and ToxB like). Structure-informed pattern searches strengthened this hypothesis by identifying MAX-effector candidates in a broad range of ascomycete phytopathogens. Strong expansion of the MAX-effector family was detected in M. oryzae and M. grisea where they seem to be particularly important since they account for 5-10% of the effector repertoire and 50% of the cloned avirulence effectors. Expression analysis indicated that the majority of M. oryzae MAX-effectors are expressed specifically during early infection suggesting important functions during biotrophic host colonization. We hypothesize that the scenario observed for MAX-effectors can serve as a paradigm for ascomycete effector diversity and that the enormous number of sequence-unrelated ascomycete effectors may in fact belong to a restricted set of structurally conserved effector families.
- INSERM U1054, Centre de Biochimie Structurale, Montpellier, France; CNRS UMR5048, Montpellier University, Montpellier, France.
Organizational Affiliation: