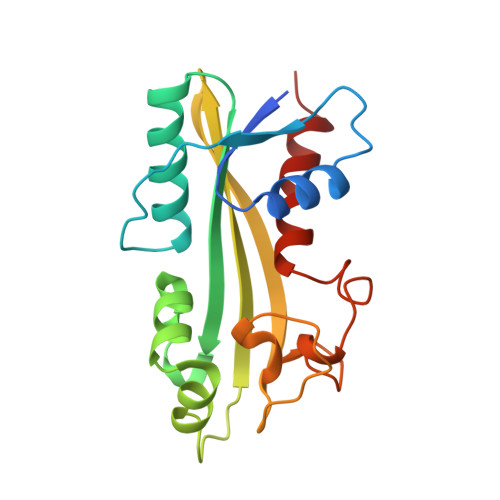
Structure-based identification of a novel NTPase from Methanococcus jannaschii.
Hwang, K.Y., Chung, J.H., Kim, S.H., Han, Y.S., Cho, Y.(1999) Nat Struct Biol 6: 691-696
- PubMed: 10404228
- DOI: https://doi.org/10.1038/10745
- Primary Citation of Related Structures:
1B78, 2MJP - PubMed Abstract:
Almost half of the entire set of predicted genomic products from Methanococcus jannaschii are classified as functionally unknown hypothetical proteins. We present a structure-based identification of the biochemical function of a protein with an as yet unknown function from a M. jannaschii gene, Mj0226. The crystal structure of Mj0226 protein determined at 2.2 A resolution reveals that the protein is a homodimer and each monomer folds into an elongated alpha/beta structure of a new fold family. Comparisons of Mj0226 protein with protein structures in the database, however, indicate that one part of the protein is homologous to some of the nucleotide-binding proteins. Biochemical analysis shows that Mj0226 protein is a novel nucleotide triphosphatase that can efficiently hydrolyze nonstandard nucleotides such as XTP to XMP or ITP to IMP, but not the standard nucleotides, in the presence of Mg2+ or Mn2+ ions.
- Structural Biology Center, Korea Institute of Science and Technology, Seoul, South Korea.
Organizational Affiliation: