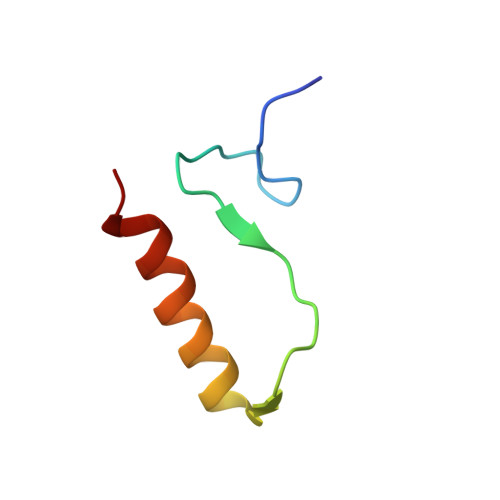
Structure and DNA-binding traits of the transition state regulator AbrB.
Olson, A.L., Tucker, A.T., Bobay, B.G., Soderblom, E.J., Moseley, M.A., Thompson, R.J., Cavanagh, J.(2014) Structure 22: 1650-1656
- PubMed: 25308864
- DOI: https://doi.org/10.1016/j.str.2014.08.018
- Primary Citation of Related Structures:
2MJG - PubMed Abstract:
The AbrB protein from Bacillus subtilis is a DNA-binding global regulator controlling the onset of a vast array of protective functions under stressful conditions. Such functions include biofilm formation, antibiotic production, competence development, extracellular enzyme production, motility, and sporulation. AbrB orthologs are known in a variety of prokaryotic organisms, most notably in all infectious strains of Clostridia, Listeria, and Bacilli. Despite its central role in bacterial response and defense, its structure has been elusive because of its highly dynamic character. Orienting its N- and C-terminal domains with respect to one another has been especially problematic. Here, we have generated a structure of full-length, tetrameric AbrB using nuclear magnetic resonance, chemical crosslinking, and mass spectrometry. We note that AbrB possesses a strip of positive electrostatic potential encompassing its DNA-binding region and that its C-terminal domain aids in DNA binding.
- Department of Molecular and Structural Biochemistry, North Carolina State University, 128 Polk Hall, Raleigh, NC 27659, USA.
Organizational Affiliation: