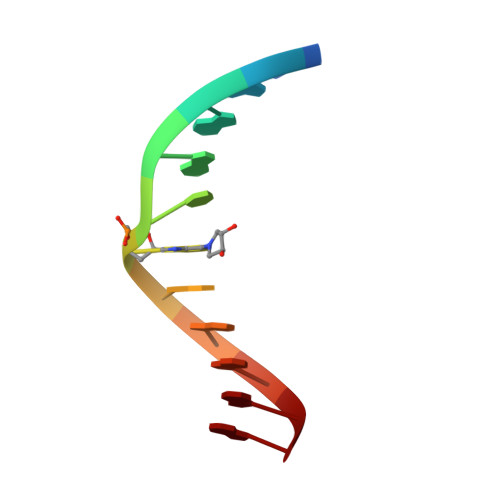
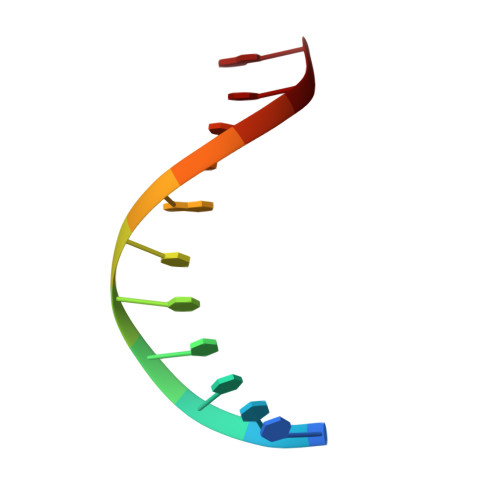
Structures of Exocyclic R,R- and S,S-N(6),N(6)-(2,3-Dihydroxybutan-1,4-diyl)-2'-Deoxyadenosine Adducts Induced by 1,2,3,4-Diepoxybutane.
Kowal, E.A., Seneviratne, U., Wickramaratne, S., Doherty, K.E., Cao, X., Tretyakova, N., Stone, M.P.(2014) Chem Res Toxicol 27: 805-817
- PubMed: 24741991 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/tx400472p
- Primary Citation Related Structures:
2MHX, 2MHZ - PubMed Abstract:
1,3-Butadiene (BD) is an industrial and environmental chemical present in urban air and cigarette smoke, and is classified as a human carcinogen. It is oxidized by cytochrome P450 to form 1,2,3,4-diepoxybutane (DEB); DEB bis-alkylates the N(6) position of adenine in DNA. Two enantiomers of bis-N(6)-dA adducts of DEB have been identified: R,R-N(6),N(6)-(2,3-dihydroxybutan-1,4-diyl)-2'-deoxyadenosine (R,R-DHB-dA), and S,S-N(6),N(6)-(2,3-dihydroxybutan-1,4-diyl)-2'-deoxyadenosine (S,S-DHB-dA) [ Seneviratne , U. , Antsypovich , S. , Dorr , D. Q. , Dissanayake , T. , Kotapati , S. , and Tretyakova , N. ( 2010 ) Chem. Res. Toxicol. 23 , 1556 -1567 ]. Herein, the R,R-DHB-dA and S,S-DHB-dA adducts have been incorporated into the 5'-d(C(1)G(2)G(3)A(4)C(5)X(6)A(7)G(8)A(9)A(10)G(11))-3':5'-d(C(12)T(13)T(14)C(15)T(16)T(17)G(18)T(19)C(20)C(21)G(22))-3' duplex [X(6) = R,R-DHB-dA (R(6)) or S,S-DHB-dA (S(6))]. The structures of the duplexes were determined by molecular dynamics calculations, which were restrained by experimental distances obtained from NMR data. Both the R,R- and S,S-DHB-dA adducts are positioned in the major groove of DNA. In both instances, the bulky 3,4-dihydroxypyrrolidine rings are accommodated by an out-of-plane rotation about the C6-N(6) bond of the bis-alkylated adenine. In both instances, the directionality of the dihydroxypyrrolidine ring is evidenced by the pattern of NOEs between the 3,4-dihydroxypyrrolidine protons and DNA. Also in both instances, the anti conformation of the glycosyl bond is maintained, which combined with the out-of-plane rotation about the C6-N(6) bond, allows the complementary thymine, T(17), to remain stacked within the duplex, and form one hydrogen bond with the modified base, between the imine nitrogen of the modified base and the T(17) N3H imino proton. The loss of the second Watson-Crick hydrogen bonding interaction at the lesion sites correlates with the lower thermal stabilities of the R,R- and S,S-DHB-dA duplexes, as compared to the corresponding unmodified duplex. The reduced base stacking at the adduct sites may also contribute to the thermal instability.
- Department of Chemistry, Center in Molecular Toxicology, and Center for Structural Biology, Stevenson Science Center, Vanderbilt University , 2201 West End Avenue, Nashville, Tennessee 37235, United States.
Organizational Affiliation: