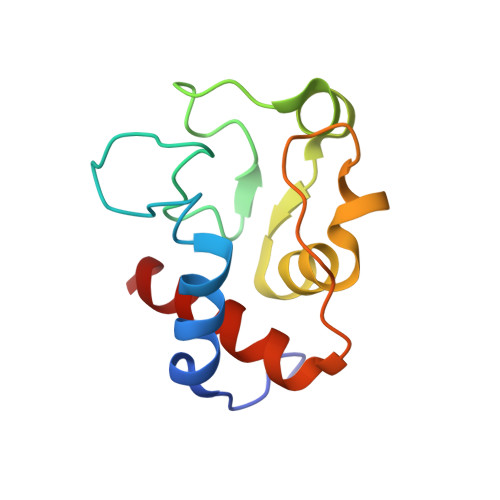
Structural Basis for Cytochrome c Y67H Mutant to Function as a Peroxidase
Lan, W.X., Wang, Z.H., Yang, Z.Z., Ying, T.L., Zhang, X., Tan, X.S., Liu, M., Cao, C.Y., Huang, Z.X.(2014) PLoS One 9: e107305-e107305r
- PubMed: 25210769
- DOI: https://doi.org/10.1371/journal.pone.0107305
- Primary Citation Related Structures:
2MHM - PubMed Abstract:
The catalytic activity of cytochrome c (cyt c) to peroxidize cardiolipin to its oxidized form is required for the release of pro-apoptotic factors from mitochondria, and for execution of the subsequent apoptotic steps. However, the structural basis for this peroxidation reaction remains unclear. In this paper, we determined the three-dimensional NMR solution structure of yeast cyt c Y67H variant with high peroxidase activity, which is almost similar to that of its native form. The structure reveals that the hydrogen bond between Met80 and residue 67 is disrupted. This change destabilizes the sixth coordination bond between heme Fe(3+) ion and Met80 sulfur atom in the Y67H variant, and further makes it more easily be broken at low pH conditions. The steady-state studies indicate that the Y67H variant has the highest peroxidase activities when pH condition is between 4.0 and 5.2. Finally, a mechanism is suggested for the peroxidation of cardiolipin catalyzed by the Y67H variant, where the residue His67 acts as a distal histidine, its protonation facilitates O-O bond cleavage of H2O2 by functioning as an acidic catalyst.
- State Key Laboratory of Natural Products and Bioorganic Chemistry, Shanghai Institute of Organic Chemistry, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: