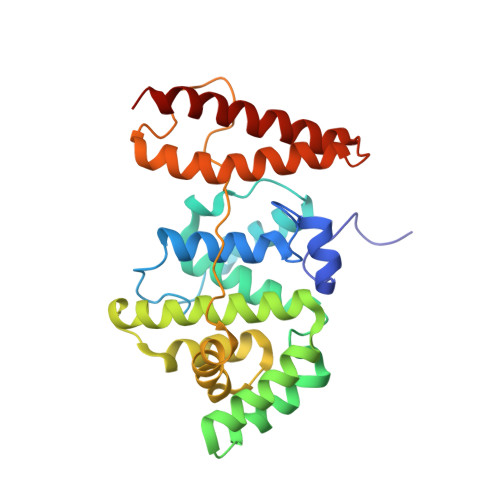
The Structure of the RLIP76 RhoGAP-Ral Binding Domain Dyad: Fixed Position of the Domains Leads to Dual Engagement of Small G Proteins at the Membrane.
Rajasekar, K.V., Campbell, L.J., Nietlispach, D., Owen, D., Mott, H.R.(2013) Structure 21: 2131-2142
- PubMed: 24207123 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2013.09.007
- Primary Citation Related Structures:
2MBG - PubMed Abstract:
RLIP76 is an effector for Ral small GTPases, which in turn lie downstream of the master regulator Ras. Evidence is growing that Ral and RLIP76 play a role in tumorigenesis, invasion, and metastasis. RLIP76 contains both a RhoGAP domain and a Ral binding domain (GBD) and is, therefore, a node between Ras and Rho family signaling. The structure of the RhoGAP-GBD dyad reveals that the RLIP76 RhoGAP domain adopts a canonical RhoGAP domain structure and that the linker between the two RLIP76 domains is structured, fixing the orientation of the two domains and allowing RLIP76 to interact with Rho-family GTPases and Ral simultaneously. However, the juxtaposed domains do not influence each other functionally, suggesting that the RLIP76-Ral interaction controls cellular localization and that the fixed orientation of the two domains orientates the RhoGAP domain with respect to the membrane, allowing it to be perfectly poised to engage its target G proteins.
- Department of Biochemistry, University of Cambridge, 80 Tennis Court Road, Cambridge CB2 1GA, UK.
Organizational Affiliation: