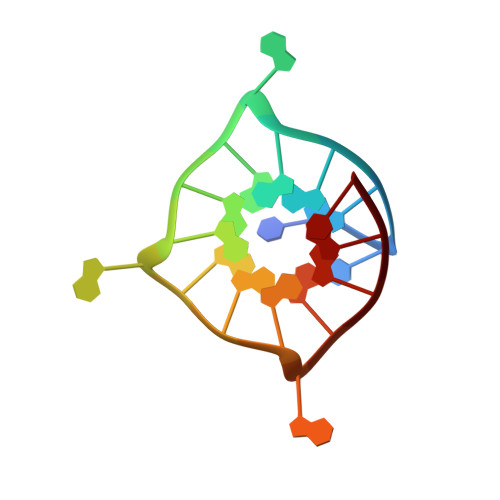
Formation of g-quadruplexes in poly-g sequences: structure of a propeller-type parallel-stranded g-quadruplex formed by a g15 stretch.
Sengar, A., Heddi, B., Phan, A.T.(2014) Biochemistry 53: 7718-7723
- PubMed: 25375976 Search on PubMed
- DOI: https://doi.org/10.1021/bi500990v
- Primary Citation Related Structures:
2MB2 - PubMed Abstract:
Poly-G sequences are found in different genomes including human and have the potential to form higher-order structures with various applications. Previously, long poly-G sequences were thought to lead to multiple possible ways of G-quadruplex folding, rendering their structural characterization challenging. Here we investigate the structure of G-quadruplexes formed by poly-G sequences d(TTG(n)T), where n = 12 to 19. Our data show the presence of multiple and/or higher-order G-quadruplex structures in most sequences. Strikingly, NMR spectra of the TTG₁₅T sequence containing a stretch of 15 continuous guanines are exceptionally well-resolved and indicate the formation of a well-defined G-quadruplex structure. The NMR solution structure of this sequence revealed a propeller-type parallel-stranded G-quadruplex containing three G-tetrad layers and three single-guanine propeller loops. The same structure can potentially form anywhere along a long G(n) stretch, making it unique for molecular recognition by other cellular molecules.
- School of Physical and Mathematical Sciences, Nanyang Technological University , Singapore 637371, Singapore.
Organizational Affiliation: