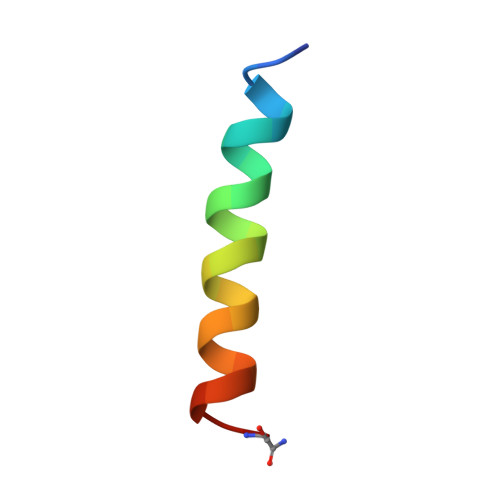
Two-dimensional 1H NMR experiments show that the 23-residue magainin antibiotic peptide is an alpha-helix in dodecylphosphocholine micelles, sodium dodecylsulfate micelles, and trifluoroethanol/water solution.
Gesell, J., Zasloff, M., Opella, S.J.(1997) J Biomol NMR 9: 127-135
- PubMed: 9090128 Search on PubMed
- DOI: https://doi.org/10.1023/a:1018698002314
- Primary Citation Related Structures:
2MAG - PubMed Abstract:
Magainin2 is a 23-residue antibiotic peptide that disrupts the ionic gradient across certain cell membranes. Two-dimensional 1H NMR spectroscopy was used to investigate the structure of the peptide in three of the membrane environments most commonly employed in biophysical studies. Sequence-specific resonance assignments were determined for the peptide in perdeuterated dodecylphosphocholine (DPC) and sodium dodecylsulfate micelles and confirmed for the peptide in 2,2,2-trifluoroethanol solution. The secondary structure is shown to be helical in all of the solvent systems. The NMR data were used as a set of restraints for a simulated annealing protocol that generated a family of three-dimensional structures of the peptide in DPC micelles, which superimposed best between residues 4 and 20. For these residues, the mean pairwise rms difference for the backbone atoms is 0.47 +/- 0.10 A from the average structure. The calculated peptide structures appear to be curved, with the bend centered at residues Phe12 and Gly13.
- Department of Chemistry, University of Pennsylvania, PA 19104, USA.
Organizational Affiliation: