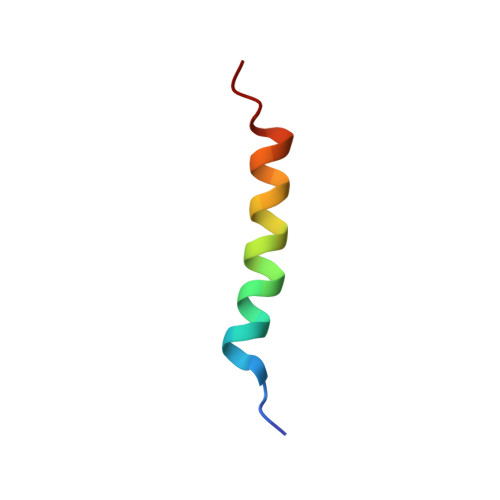
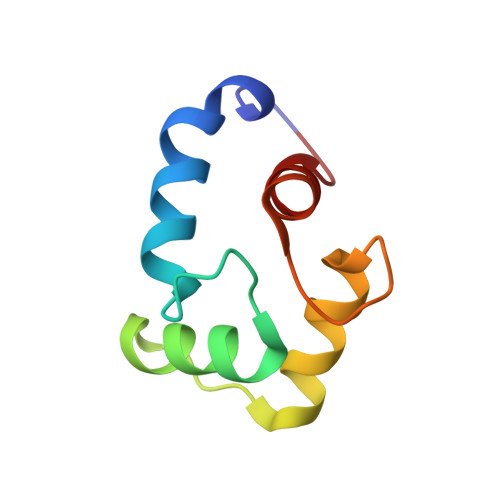
Calcium triggers reversal of calmodulin on nested anti-parallel sites in the IQ motif of the neuronal voltage-dependent sodium channel NaV1.2.
Hovey, L., Fowler, C.A., Mahling, R., Lin, Z., Miller, M.S., Marx, D.C., Yoder, J.B., Kim, E.H., Tefft, K.M., Waite, B.C., Feldkamp, M.D., Yu, L., Shea, M.A.(2017) Biophys Chem 224: 1-19
- PubMed: 28343066 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bpc.2017.02.006
- Primary Citation Related Structures:
2M5E - PubMed Abstract:
Several members of the voltage-gated sodium channel family are regulated by calmodulin (CaM) and ionic calcium. The neuronal voltage-gated sodium channel Na V 1.2 contains binding sites for both apo (calcium-depleted) and calcium-saturated CaM. We have determined equilibrium dissociation constants for rat Na V 1.2 IQ motif [IQRAYRRYLLK] binding to apo CaM (~3nM) and (Ca 2+ ) 4 -CaM (~85nM), showing that apo CaM binding is favored by 30-fold. For both apo and (Ca 2+ ) 4 -CaM, NMR demonstrated that Na V 1.2 IQ motif peptide (Na V 1.2 IQp ) exclusively made contacts with C-domain residues of CaM (CaM C ). To understand how calcium triggers conformational change at the CaM-IQ interface, we determined a solution structure (2M5E.pdb) of (Ca 2+ ) 2 -CaM C bound to Na V 1.2 IQp . The polarity of (Ca 2+ ) 2 -CaM C relative to the IQ motif was opposite to that seen in apo CaM C -Na v 1.2 IQp (2KXW), revealing that CaM C recognizes nested, anti-parallel sites in Na v 1.2 IQp . Reversal of CaM may require transient release from the IQ motif during calcium binding, and facilitate a re-orientation of CaM N allowing interactions with non-IQ Na V 1.2 residues or auxiliary regulatory proteins interacting in the vicinity of the IQ motif.
- Department of Biochemistry, University of Iowa, 52242-1109 Iowa City, United States.
Organizational Affiliation: