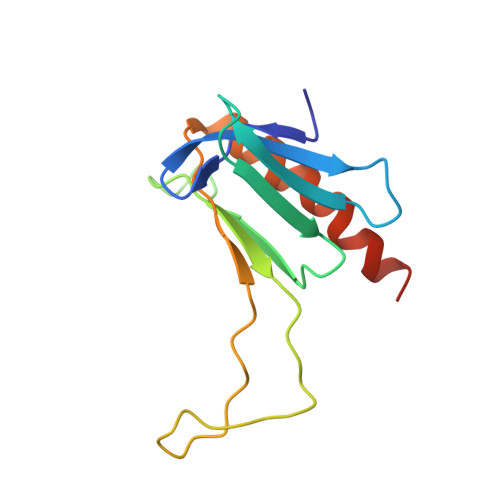
Structural and functional evidence that Rad4 competes with Rad2 for binding to the Tfb1 subunit of TFIIH in NER.
Lafrance-Vanasse, J., Arseneault, G., Cappadocia, L., Legault, P., Omichinski, J.G.(2013) Nucleic Acids Res 41: 2736-2745
- PubMed: 23295669
- DOI: https://doi.org/10.1093/nar/gks1321
- Primary Citation Related Structures:
2M14 - PubMed Abstract:
XPC/Rad4 (human/yeast) recruits transcription faction IIH (TFIIH) to the nucleotide excision repair (NER) complex through interactions with its p62/Tfb1 and XPB/Ssl2 subunits. TFIIH then recruits XPG/Rad2 through interactions with similar subunits and the two repair factors appear to be mutually exclusive within the NER complex. Here, we show that Rad4 binds the PH domain of the Tfb1 (Tfb1PH) with high affinity. Structural characterization of a Rad4-Tfb1PH complex demonstrates that the Rad4-binding interface is formed using a motif similar to one used by Rad2 to bind Tfb1PH. In vivo studies in yeast demonstrate that the N-terminal Tfb1-binding motif and C-terminal TFIIH-binding motif of Rad4 are both crucial for survival following exposure to UV irradiation. Together, these results support the hypothesis that XPG/Rad2 displaces XPC/Rad4 from the repair complex in part through interactions with the Tfb1/p62 subunit of TFIIH. The Rad4-Tfb1PH structure also provides detailed information regarding, not only the interplay of TFIIH recruitment to the NER, but also links the role of TFIIH in NER and transcription.
- Département de Biochimie, Université de Montréal C.P. 6128 Succursale Centre-Ville, Montréal, Québec, Canada H3C 3J7.
Organizational Affiliation: