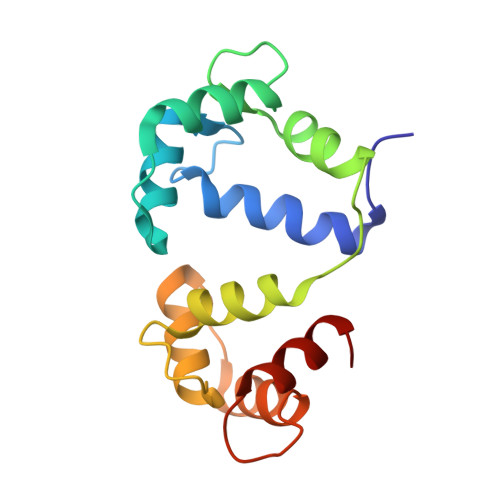
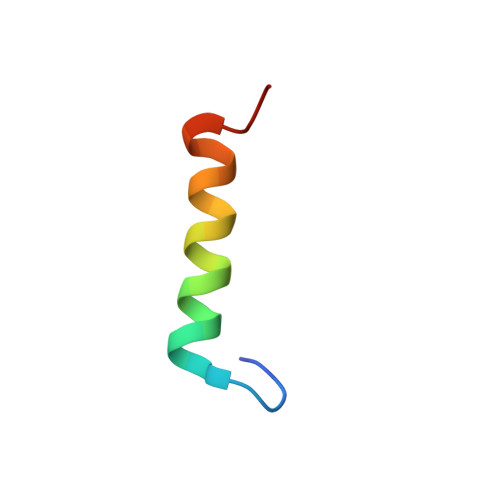
Binding orientation and specificity of calmodulin to rat olfactory cyclic nucleotide-gated ion channel.
Irene, D., Huang, J.W., Chung, T.Y., Li, F.Y., Tzen, J.T., Lin, T.H., Chyan, C.L.(2013) J Biomol Struct Dyn 31: 414-425
- PubMed: 22877078 Search on PubMed
- DOI: https://doi.org/10.1080/07391102.2012.703069
- Primary Citation Related Structures:
2M0J, 2M0K - PubMed Abstract:
Calmodulin (CaM), the primary intracellular Ca(2+) receptor, regulates a large number of key enzymes and controls a wide spectrum of important biological responses. Recognition between CaM and its target sequence in rat olfactory cyclic nucleotide-gated ion channel (OLFp) was investigated by circular dichroism (CD), fluorescence, and NMR spectroscopy. Fluorescence data showed the OLFp tightly bound to CaM with a dissociation constant of 12 nM in a 1:1 stoichiometry. Far-UV CD data showed that approximately 60% of OLFp residues formed α-helical structures when associated with CaM. NMR data showed that most of the (15)N-(1)H HSQC cross-peaks of the (15)N-labeled CaM not only shifted but also split into two sets of peaks upon association with the OLFp. Our data indicated that the two distinct CaM/OLFp complexes existed simultaneously with stable structures that were not interexchangeable within the NMR time scale. In light of the palindromic sequence of OLFp (FQRIVRLVGVIRDW) for CaM targeting, we proposed that the helical OLFp with C2 symmetry may bind to CaM in two orientations. This hypothesis is supported by the observation that only one set of (15)N-(1)H HSQC cross-peaks of the (15)N-labeled CaM was detected upon association with OLFp-M13 chimeric peptide (OLFMp), a mutated OLFp lacking the palindromic feature. The binding specificity of OLFMp to CaM was restored when the palindromic feature was destroyed. Binding modes of CaM/OLFp and CaM/OLFMp simulated by molecular docking were in accord with their distinct patterns observed in HSQC spectra. Our studies suggest that the palindromic residues in OLFp are crucial for the orientation-specific recognition by CaM.
- Department of Chemistry, National Dong Hwa University, Hualien 974, Taiwan, ROC.
Organizational Affiliation: