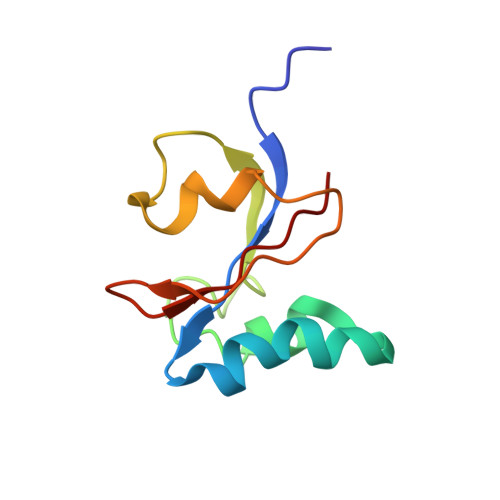
NmPin from the marine thaumarchaeote Nitrosopumilus maritimus is an active membrane associated prolyl isomerase.
Hoppstock, L., Trusch, F., Lederer, C., van West, P., Koenneke, M., Bayer, P.(2016) BMC Biol 14: 53-53
- PubMed: 27349962
- DOI: https://doi.org/10.1186/s12915-016-0274-1
- Primary Citation Related Structures:
2M08 - PubMed Abstract:
Peptidyl-prolyl isomerases (PPIases) are present in all forms of life and play a crucial role in protein folding and regulation. They catalyze the cis-trans isomerization of the peptide bond that precedes proline residues in numerous proteins. The parvulins, which is one family of PPIases, have been extensively investigated in several eukaryotes. However, nothing is known about their expression, function and localization in archaea. Here, we describe the endogenous expression, molecular structure, function and cellular localization of NmPin, a single-domain parvulin-type PPIase from Nitrosopumilus maritimus. This marine chemolithoautotrophic archaeon belongs to the globally abundant phylum Thaumarchaeota. Using high resolution NMR spectroscopy we demonstrate that the 3D structure of NmPin adopts a parvulin fold and confirmed its peptidyl-prolyl isomerase activity by protease-coupled assays and mutagenesis studies. A detailed topological analysis revealed a positively charged lysine-rich patch on the protein surface, which is conserved in all known parvulin sequences of thaumarchaeotes and targets NmPin to lipids in vitro. Immunofluorescence microscopy confirms that the protein is attached to the outer archaeal cell membrane in vivo. Transmission electron microscopy uncovered that NmPin has a uniform distribution at the membrane surface, which is correlated with a native cell shape of the prokaryote. We present a novel solution structure of a catalytically active thaumarchaeal parvulin. Our results reveal that a lysine-rich patch in NmPin mediates membrane localization. These findings provide a model whereby NmPin is located between the archaeal membrane and the surface layer and hence suggest proteins of the S-layer as the key target substrates of this parvulin.
- Department of Structural and Medicinal Biochemistry, Centre for Medical Biotechnology, University of Duisburg-Essen, Universitätsstr. 1-4, 45141, Essen, Germany.
Organizational Affiliation: