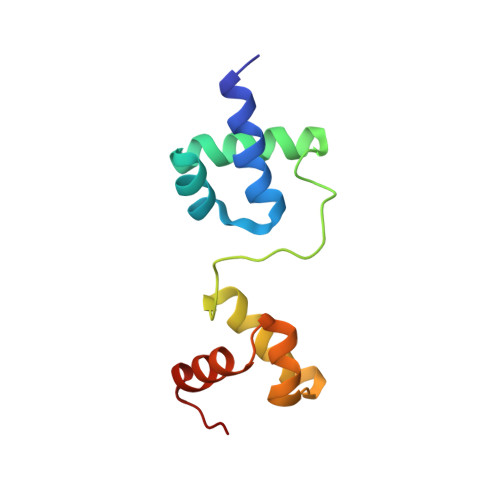
Solution properties of the archaeal CRISPR DNA repeat-binding homeodomain protein Cbp2.
Kenchappa, C.S., Heidarsson, P.O., Kragelund, B.B., Garrett, R.A., Poulsen, F.M.(2013) Nucleic Acids Res 41: 3424-3435
- PubMed: 23325851
- DOI: https://doi.org/10.1093/nar/gks1465
- Primary Citation of Related Structures:
2LVS - PubMed Abstract:
Clustered regularly interspaced short palindromic repeats (CRISPR) form the basis of diverse adaptive immune systems directed primarily against invading genetic elements of archaea and bacteria. Cbp1 of the crenarchaeal thermoacidophilic order Sulfolobales, carrying three imperfect repeats, binds specifically to CRISPR DNA repeats and has been implicated in facilitating production of long transcripts from CRISPR loci. Here, a second related class of CRISPR DNA repeat-binding protein, denoted Cbp2, is characterized that contains two imperfect repeats and is found amongst members of the crenarchaeal thermoneutrophilic order Desulfurococcales. DNA repeat-binding properties of the Hyperthermus butylicus protein Cbp2Hb were characterized and its three-dimensional structure was determined by NMR spectroscopy. The two repeats generate helix-turn-helix structures separated by a basic linker that is implicated in facilitating high affinity DNA binding of Cbp2 by tethering the two domains. Structural studies on mutant proteins provide support for Cys(7) and Cys(28) enhancing high thermal stability of Cbp2Hb through disulphide bridge formation. Consistent with their proposed CRISPR transcriptional regulatory role, Cbp2Hb and, by inference, other Cbp1 and Cbp2 proteins are closely related in structure to homeodomain proteins with linked helix-turn-helix (HTH) domains, in particular the paired domain Pax and Myb family proteins that are involved in eukaryal transcriptional regulation.
- Archaea Centre, Department of Biology, University of Copenhagen, Ole Maaløes Vej 5, DK-2200 Copenhagen, Denmark.
Organizational Affiliation: