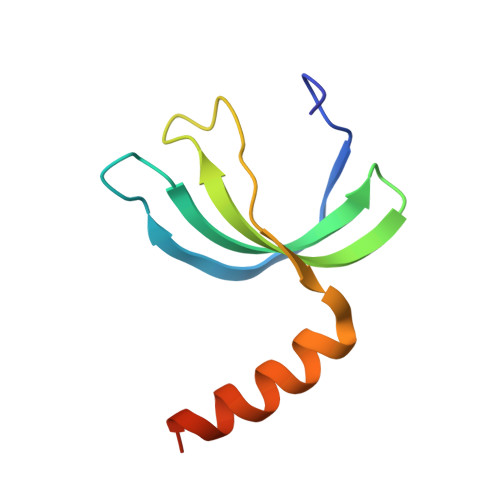
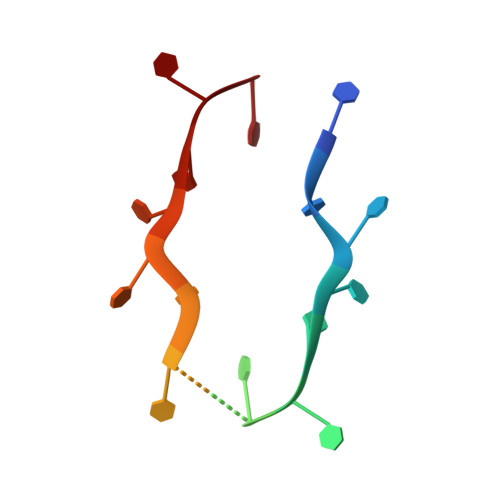
Structures of apo- and ssDNA-bound YdbC from Lactococcus lactis uncover the function of protein domain family DUF2128 and expand the single-stranded DNA-binding domain proteome.
Rossi, P., Barbieri, C.M., Aramini, J.M., Bini, E., Lee, H.W., Janjua, H., Xiao, R., Acton, T.B., Montelione, G.T.(2013) Nucleic Acids Res 41: 2756-2768
- PubMed: 23303792
- DOI: https://doi.org/10.1093/nar/gks1348
- Primary Citation Related Structures:
2LTD, 2LTT - PubMed Abstract:
Single-stranded DNA (ssDNA) binding proteins are important in basal metabolic pathways for gene transcription, recombination, DNA repair and replication in all domains of life. Their main cellular role is to stabilize melted duplex DNA and protect genomic DNA from degradation. We have uncovered the molecular function of protein domain family domain of unknown function DUF2128 (PF09901) as a novel ssDNA binding domain. This bacterial domain strongly associates into a dimer and presents a highly positively charged surface that is consistent with its function in non-specific ssDNA binding. Lactococcus lactis YdbC is a representative of DUF2128. The solution NMR structures of the 20 kDa apo-YdbC dimer and YdbC:dT(19)G(1) complex were determined. The ssDNA-binding energetics to YdbC were characterized by isothermal titration calorimetry. YdbC shows comparable nanomolar affinities for pyrimidine and mixed oligonucleotides, and the affinity is sufficiently strong to disrupt duplex DNA. In addition, YdbC binds with lower affinity to ssRNA, making it a versatile nucleic acid-binding domain. The DUF2128 family is related to the eukaryotic nuclear protein positive cofactor 4 (PC4) family and to the PUR family both by fold similarity and molecular function.
- Department of Molecular Biology and Biochemistry, Center for Advanced Biotechnology and Medicine, and the Northeast Structural Genomics Consortium, Rutgers, The State University of New Jersey, Piscataway, NJ 08854, USA.
Organizational Affiliation: