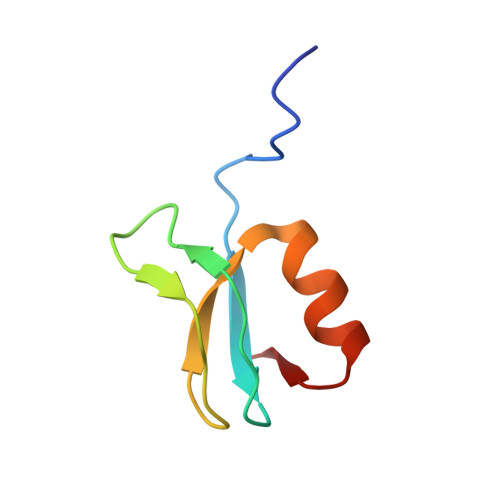
Structural and biochemical characterization of phage lambda FI protein (gpFI) reveals a novel mechanism of DNA packaging chaperone activity.
Popovic, A., Wu, B., Arrowsmith, C.H., Edwards, A.M., Davidson, A.R., Maxwell, K.L.(2012) J Biological Chem 287: 32085-32095
- PubMed: 22801427
- DOI: https://doi.org/10.1074/jbc.M112.378349
- Primary Citation of Related Structures:
2LSM - PubMed Abstract:
One of the final steps in the morphogenetic pathway of phage λ is the packaging of a single genome into a preformed empty head structure. In addition to the terminase enzyme, the packaging chaperone, FI protein (gpFI), is required for efficient DNA packaging. In this study, we demonstrate an interaction between gpFI and the major head protein, gpE. Amino acid substitutions in gpFI that reduced the strength of this interaction also decreased the biological activity of gpFI, implying that this head binding activity is essential for the function of gpFI. We also show that gpFI is a two-domain protein, and the C-terminal domain is responsible for the head binding activity. Using nuclear magnetic resonance spectroscopy, we determined the three-dimensional structure of the C-terminal domain and characterized the helical nature of the N-terminal domain. Through structural comparisons, we were able to identify two previously unannotated prophage-encoded proteins with tertiary structures similar to gpFI, although they lack significant pairwise sequence identity. Sequence analysis of these diverse homologues led us to identify related proteins in a variety of myo- and siphophages, revealing that gpFI function has a more highly conserved role in phage morphogenesis than was previously appreciated. Finally, we present a novel model for the mechanism of gpFI chaperone activity in the DNA packaging reaction of phage λ.
- Department of Biochemistry, University of Toronto, Toronto, Ontario M5S 1A8, Canada.
Organizational Affiliation: