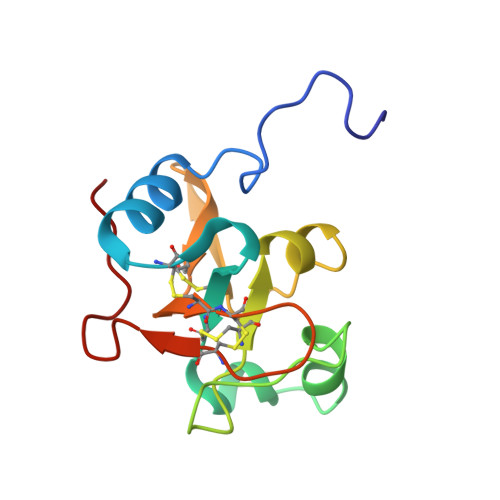
Analysis of the structure and conformational states of DewA gives insight into the assembly of the fungal hydrophobins
Morris, V.K., Kwan, A.H., Sunde, M.(2012) J Mol Biology 6: 83-86
- PubMed: 23137797
- DOI: https://doi.org/10.1016/j.jmb.2012.10.021
- Primary Citation Related Structures:
2LSH - PubMed Abstract:
The hydrophobin DewA from the fungus Aspergillus nidulans is a highly surface-active protein that spontaneously self-assembles into amphipathic monolayers at hydrophobic:hydrophilic interfaces. These monolayers are composed of fibrils that are a form of functional amyloid. While there has been significant interest in the use of DewA for a variety of surface coatings and as an emulsifier in biotechnological applications, little is understood about the structure of the protein or the mechanism of self-assembly. We have solved the solution NMR structure of DewA. While the pattern of four disulfide bonds that is a defining feature of hydrophobins is conserved, the arrangement and composition of secondary-structure elements in DewA are quite different to what has been observed in other hydrophobin structures. In addition, we demonstrate that DewA populates two conformations in solution, both of which are assembly competent. One conformer forms a dimer at high concentrations, but this dimer is off-pathway to fibril formation and may represent an assembly control mechanism. These data highlight the structural differences between fibril-forming hydrophobins and those that form amorphous monolayers. This work will open up new opportunities for the engineering of hydrophobins with novel biotechnological applications.
- School of Molecular Bioscience, University of Sydney, NSW 2006, Australia.
Organizational Affiliation: