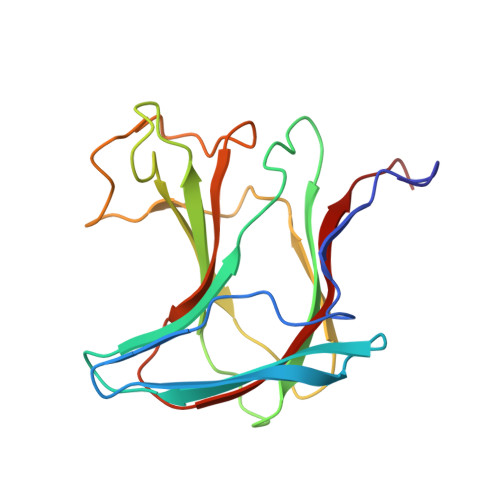
Solution structure, dynamics and binding studies of a family 11 carbohydrate-binding module from Clostridium thermocellum (CtCBM11).
Viegas, A., Sardinha, J., Freire, F., Duarte, D.F., Carvalho, A.L., Fontes, C.M., Romao, M.J., Macedo, A.L., Cabrita, E.J.(2013) Biochem J 451: 289-300
- PubMed: 23356867 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20120627
- Primary Citation Related Structures:
2LRO, 2LRP - PubMed Abstract:
Non-catalytic cellulosomal CBMs (carbohydrate-binding modules) are responsible for increasing the catalytic efficiency of cellulosic enzymes by selectively putting the substrate (a wide range of poly- and oligo-saccharides) and enzyme into close contact. In the present study we carried out an atomistic rationalization of the molecular determinants of ligand specificity for a family 11 CBM from thermophilic Clostridium thermocellum [CtCBM11 (C. thermocellum CBM11)], based on a NMR and molecular modelling approach. We have determined the NMR solution structure of CtCBM11 at 25°C and 50°C and derived information on the residues of the protein that are involved in ligand recognition and on the influence of the length of the saccharide chain on binding. We obtained models of the CtCBM11-cellohexaose and CtCBM11-cellotetraose complexes by docking in accordance with the NMR experimental data. Specific ligand-protein CH-π and Van der Waals interactions were found to be determinant for the stability of the complexes and for defining specificity. Using the order parameters derived from backbone dynamics analysis in the presence and absence of ligand and at 25°C and 50°C, we determined that the protein's backbone conformational entropy is slightly positive. This data in combination with the negative binding entropy calculated from ITC (isothermal titration calorimetry) studies supports a selection mechanism where a rigid protein selects a defined oligosaccharide conformation.
- REQUIMTE-CQFB, Dep. de Química, Faculdade de Ciências e Tecnologia, Universidade Nova de Lisboa, 2829-516 Caparica, Portugal.
Organizational Affiliation: