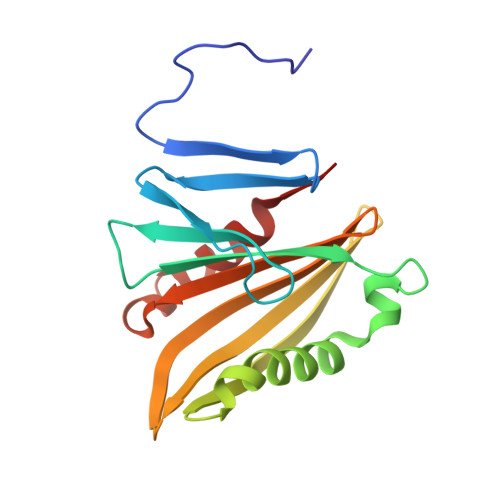
Solution structure of CyanoP from Synechocystis sp. PCC 6803: New insights on the structural basis for functional specialization amongst PsbP family proteins
Jackson, S.A., Hinds, M.G., Eaton-Rye, J.J.(2012) Biochim Biophys Acta
- PubMed: 22414666
- DOI: https://doi.org/10.1016/j.bbabio.2012.02.032
- Primary Citation Related Structures:
2LNJ - PubMed Abstract:
The structure of the CyanoP subunit of photosystem II from the cyanobacterium Synechocystis sp. PCC 6803 has been determined in solution by Nuclear Magnetic Resonance spectroscopy. Combined with homology modeling of PsbP-like structures we have identified distinct structural differences between PsbP homologues which may account for the functional differences apparent between members of this protein family. A surface cleft containing a large number of conserved residues found only in CyanoP and PsbP-like homologues has been identified and our findings suggest that one of the potential cation binding sites found in CyanoP may be functionally significant. Evidence for the evolution and divergence of the PsbP super family is presented from a structural perspective including identification of residues which distinguish the PsbP family from unrelated proteins with a similar domain fold. This article is part of a Special Issue entitled: Photosynthesis Research for Sustainability: from Natural to Artificial.
- Department of Biochemistry, University of Otago, Dunedin, New Zealand.
Organizational Affiliation: