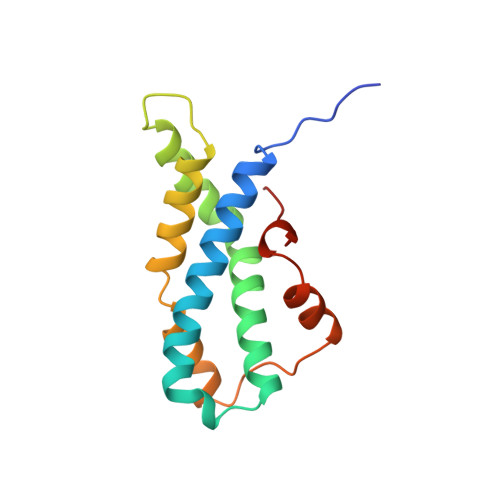
1.35 and 2.07 A resolution structures of the red abalone sperm lysin monomer and dimer reveal features involved in receptor binding.
Kresge, N., Vacquier, V.D., Stout, C.D.(2000) Acta Crystallogr D Biol Crystallogr 56: 34-41
- PubMed: 10666624
- DOI: https://doi.org/10.1107/s0907444999014626
- Primary Citation Related Structures:
2LIS, 2LYN - PubMed Abstract:
Abalone sperm use lysin to make a hole in the egg's protective vitelline envelope (VE). When released from sperm, lysin first binds to the VE receptor for lysin (VERL) then dissolves the VE by a non-enzymatic mechanism. The structures of the monomeric and dimeric forms of Haliotis rufescens (red abalone) lysin, previously solved at 1.90 and 2.75 A, respectively, have now been refined to 1.35 and 2.07 A, respectively. The monomeric form of lysin was refined using previously obtained crystallization conditions, while the dimer was solved in a new crystal form with four molecules (two dimers) per asymmetric unit. These high-resolution structures reveal alternate residue conformations, enabling a thorough analysis of the conserved residues contributing to the amphipathic nature of lysin. The availability of five independent high-resolution copies of lysin permits comparisons leading to insights on the local flexibility of lysin and alternative conformations of the hypervariable N-terminus, thought to be involved in species-specific receptor recognition. The new analysis led to the discovery of the basic nature of a cleft formed upon dimerization and a patch of basic residues in the dimer interface. Identification of these features was not possible at lower resolution. In light of this new information, a model explaining the binding of sperm lysin to egg VERL and the subsequent dissolution of the egg VE is proposed.
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA 92037-1093, USA.
Organizational Affiliation: