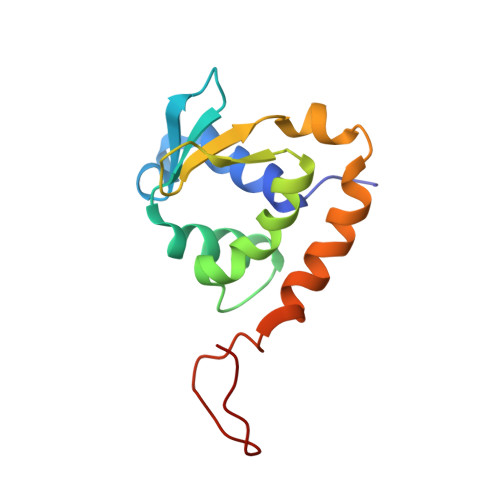
Autoinhibition of ETV6 (TEL) DNA Binding: Appended Helices Sterically Block the ETS Domain.
Coyne, H.J., De, S., Okon, M., Green, S.M., Bhachech, N., Graves, B.J., McIntosh, L.P.(2012) J Mol Biology 421: 67-84
- PubMed: 22584210 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2012.05.010
- Primary Citation Related Structures:
2LF7, 2LF8 - PubMed Abstract:
ETV6 (or TEL), a transcriptional repressor belonging to the ETS family, is frequently involved in chromosomal translocations linked with human cancers. It displays a DNA-binding mode distinct from other ETS proteins due to the presence of a self-associating PNT domain. In this study, we used NMR spectroscopy to dissect the structural and dynamic bases for the autoinhibition of ETV6 DNA binding by sequences C-terminal to its ETS domain. The C-terminal inhibitory domain (CID) contains two helices, H4 and H5, which sterically block the DNA-binding interface of the ETS domain. Importantly, these appended helices are only marginally stable as revealed by amide hydrogen exchange and (15)N relaxation measurements. The CID is thus poised to undergo a facile conformational change as required for DNA binding. The CID also dampens millisecond timescale motions of the ETS domain hypothesized to be critical for the recognition of specific ETS target sequences. This work illustrates the use of appended sequences on conserved structural domains to generate biological diversity and complements previous studies of the allosteric mechanism of ETS1 autoinhibition to reveal both common and divergent features underlying the regulation of DNA binding by ETS transcription factors.
- Department of Biochemistry and Molecular Biology, University of British Columbia, Vancouver, BC, Canada V6T 1Z3.
Organizational Affiliation: