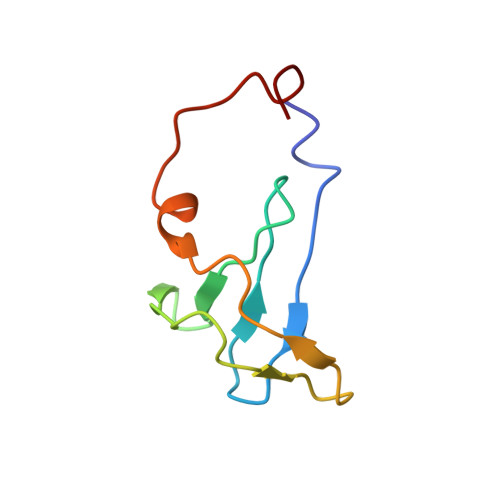
Molecular basis of the cooperative binding of Cu(I) and Cu(II) to the CopK protein from Cupriavidus metallidurans CH34.
Ash, M.R., Chong, L.X., Maher, M.J., Hinds, M.G., Xiao, Z., Wedd, A.G.(2011) Biochemistry 50: 9237-9247
- PubMed: 21936507 Search on PubMed
- DOI: https://doi.org/10.1021/bi200841f
- Primary Citation Related Structures:
2LEL - PubMed Abstract:
The bacterium Cupriavidus metallidurans CH34 is resistant to high environmental concentrations of many metal ions. Upon copper challenge, it upregulates the periplasmic protein CopK (8.3 kDa). The function of CopK in the copper resistance response is ill-defined, but CopK demonstrates an intriguing cooperativity: occupation of a high-affinity Cu(I) binding site generates a high-affinity Cu(II) binding site, and the high-affinity Cu(II) binding enhances Cu(I) binding. Native CopK and targeted variants were examined by chromatographic, spectroscopic, and X-ray crystallographic probes. Structures of two distinct forms of Cu(I)Cu(II)-CopK were defined, and structural changes associated with occupation of the Cu(II) site were demonstrated. In solution, monomeric Cu(I)Cu(II)-CopK features the previously elucidated Cu(I) site in Cu(I)-CopK, formed from four S(δ) atoms of Met28, -38, -44, and -54 (site 4S). Binding of Cu(I) to apo-CopK induces a conformational change that releases the C-terminal β-strand from the β-sandwich structure. In turn, this allows His70 and N-terminal residues to form a large loop that includes the Cu(II) binding site. In crystals, a polymeric form of Cu(I)Cu(II)-CopK displays a Cu(I) site defined by the S(δ) atoms of Met26, -38, and -54 (site 3S) and an exogenous ligand (modeled as H(2)O) and a Cu(II) site that bridges dimeric CopK molecules. The 3S Cu(I) binding mode observed in crystals was demonstrated in solution in protein variant M44L where site 4S is disabled. The intriguing copper binding chemistry of CopK provides molecular insight into Cu(I) transfer processes. The adaptable nature of the Cu(I) coordination sphere in methionine-rich clusters allows copper to be relayed between clusters during transport across membranes in molecular pumps such as CusA and Ctr1.
- School of Molecular Bioscience, University of Sydney, NSW 2006, Australia.
Organizational Affiliation: