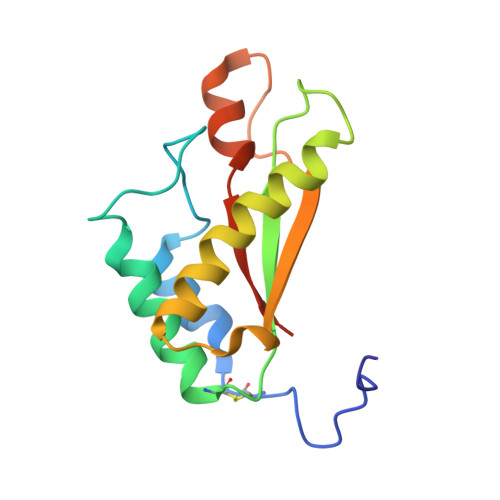
Molecular Structure and Peptidoglycan Recognition of Mycobacterium tuberculosis ArfA (Rv0899).
Yao, Y., Barghava, N., Kim, J., Niederweis, M., Marassi, F.M.(2012) J Mol Biology 416: 208-220
- PubMed: 22206986 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.12.030
- Primary Citation Related Structures:
2LBT, 2LCA - PubMed Abstract:
Mycobacterium tuberculosis ArfA (Rv0899) is a membrane protein encoded by an operon that is required for supporting bacterial growth in acidic environments. Its C-terminal domain (C domain) shares significant sequence homology with the OmpA-like family of peptidoglycan-binding domains, suggesting that its physiological function in acid stress protection may be related to its interaction with the mycobacterial cell wall. Previously, we showed that ArfA forms three independently structured modules, and we reported the structure of its central domain (B domain). Here, we describe the high-resolution structure and dynamics of the C domain, we identify ArfA as a peptidoglycan-binding protein and we elucidate the molecular basis for its specific recognition of diaminopimelate-type peptidoglycan. The C domain of ArfA adopts the characteristic fold of the OmpA-like family. It exhibits pH-dependent conformational dynamics (with significant heterogeneity at neutral pH and a more ordered structure at acidic pH), which could be related to its acid stress response. The C domain associates tightly with polymeric peptidoglycan isolated from M. tuberculosis and also associates with a soluble peptide intermediate of peptidoglycan biosynthesis. This enabled us to characterize the peptidoglycan binding site where five highly conserved ArfA residues, including two key arginines, establish the specificity for diaminopimelate- but not Lys-type peptidoglycan. ArfA is the first peptidoglycan-binding protein to be identified in M. tuberculosis. Its functions in acid stress protection and peptidoglycan binding suggest a link between the acid stress response and the physicochemical properties of the mycobacterial cell wall.
- Sanford Burnham Medical Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: