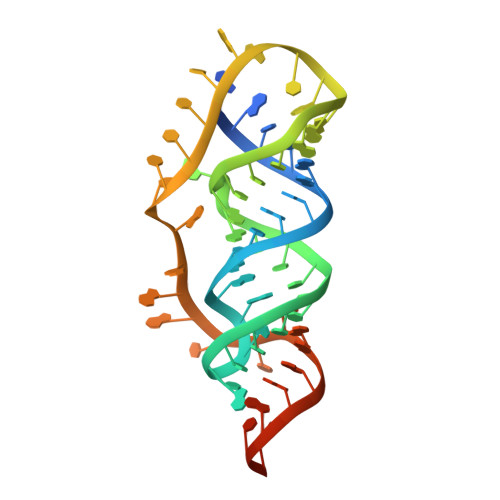
An equilibrium-dependent retroviral mRNA switch regulates translational recoding
Houck-Loomis, B., Durney, M.A., Salguero, C., Shankar, N., Nagle, J.M., Goff, S.P., D Souza, V.M.(2011) Nature 480: 561-564
- PubMed: 22121021 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature10657
- Primary Citation Related Structures:
2LC8 - PubMed Abstract:
Most retroviruses require translational recoding of a viral messenger RNA stop codon to maintain a precise ratio of structural (Gag) and enzymatic (Pol) proteins during virus assembly. Pol is expressed exclusively as a Gag-Pol fusion either by ribosomal frameshifting or by read-through of the gag stop codon. Both of these mechanisms occur infrequently and only affect 5-10% of translating ribosomes, allowing the virus to maintain the critical Gag to Gag-Pol ratio. Although it is understood that the frequency of the recoding event is regulated by cis RNA motifs, no mechanistic explanation is currently available for how the critical protein ratio is maintained. Here we present the NMR structure of the murine leukaemia virus recoding signal and show that a protonation-dependent switch occurs to induce the active conformation. The equilibrium is such that at physiological pH the active, read-through permissive conformation is populated at approximately 6%: a level that correlates with in vivo protein quantities. The RNA functions by a highly sensitive, chemo-mechanical coupling tuned to ensure an optimal read-through frequency. Similar observations for a frameshifting signal indicate that this novel equilibrium-based mechanism may have a general role in translational recoding.
- Department of Biochemistry and Molecular Biophysics, Howard Hughes Medical Institute, Columbia University, New York, New York 10032, USA.
Organizational Affiliation: