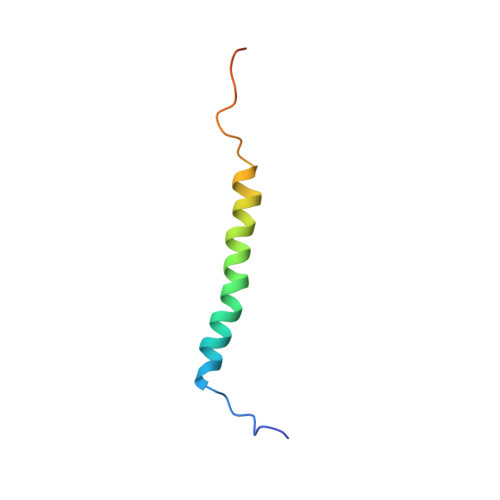
Integrin Alpha1 Has a Long Helix, Extending from the Transmembrane Region to the Cytoplasmic Tail in Detergent Micelles
Lai, C., Liu, X., Tian, C., Wu, F.(2013) PLoS One 8: e62954-e62954
- PubMed: 23646163
- DOI: https://doi.org/10.1371/journal.pone.0062954
- Primary Citation Related Structures:
2L8S - PubMed Abstract:
Integrin proteins are very important adhesion receptors that mediate cell-cell and cell-extracellular matrix interactions. They play essential roles in cell signaling and the regulation of cellular shape, motility, and the cell cycle. Here, the transmembrane and cytoplasmic (TMC) domains of integrin α1 and β1 were over-expressed and purified in detergent micelles. The structure and backbone relaxations of α1-TMC in LDAO micelles were determined and analyzed using solution NMR. A long helix, extending from the transmembrane region to the cytoplasmic tail, was observed in α1-TMC. Structural comparisons of α1-TMC with reported αIIb-TMC domains indicated different conformations in the transmembrane regions and cytoplasmic tails. An NMR titration experiment indicated weak interactions between α1-TMC and β1-TMC through several α1-TMC residues located at its N-terminal juxta-transmembrane region and C-terminal extended helix region.
- Hefei National Laboratory for Physical Science at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, China.
Organizational Affiliation: