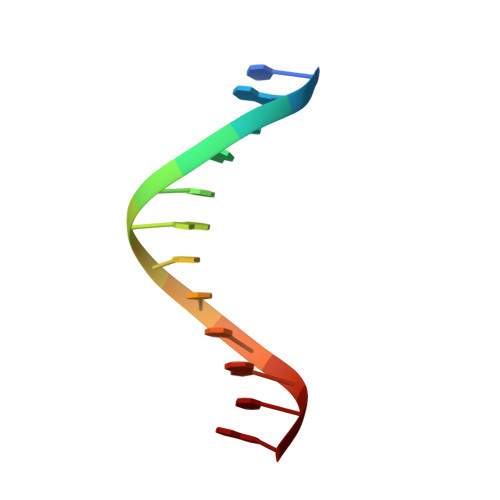
Solution structure of the Dickerson DNA dodecamer containing a single ribonucleotide.
DeRose, E.F., Perera, L., Murray, M.S., Kunkel, T.A., London, R.E.(2012) Biochemistry 51: 2407-2416
- PubMed: 22390730
- DOI: https://doi.org/10.1021/bi201710q
- Primary Citation Related Structures:
2L7D - PubMed Abstract:
Ribonucleotides are frequently incorporated into DNA during replication. They are recognized and processed by several cellular enzymes, and their continued presence in the yeast nuclear genome results in replicative stress and genome instability. Thus, it is important to understand the effects of isolated ribonucleotide incorporation on DNA structure. With this goal in mind, we describe the nuclear magnetic resonance structure of the self-complementary Dickerson dodecamer sequence [d(CGC)rGd(AATTCGCG)](2) containing two symmetrically positioned riboguanosines. The absence of an observable H(1)-H(2) scalar coupling interaction indicates a C3'-endo conformation for the ribose. Longer-range structural perturbations resulting from the presence of the ribonucleotide are limited to the adjacent and transhelical nucleotides, while the global B-form DNA structure is maintained. Because crystallographic studies have indicated that isolated ribonucleotides promote global B → A transitions, we also performed molecular modeling analyses to evaluate the structural consequences of higher ribonucleotide substitution levels. Increasing the ribonucleotide content increased the minor groove width toward values more similar to that of A-DNA, but even 50% ribonucleotide substitution did not fully convert the B-DNA to A-DNA. Comparing our structure with the structure of an RNase H2-bound DNA supports the conclusion that, as with other DNA-protein complexes, the DNA conformation is strongly influenced by the interaction with the protein.
- Laboratory of Structural Biology, National Institute of Environmental Health Sciences, National Institutes of Health, Department of Health and Human Services, Research Triangle Park, North Carolina 27709, United States.
Organizational Affiliation: