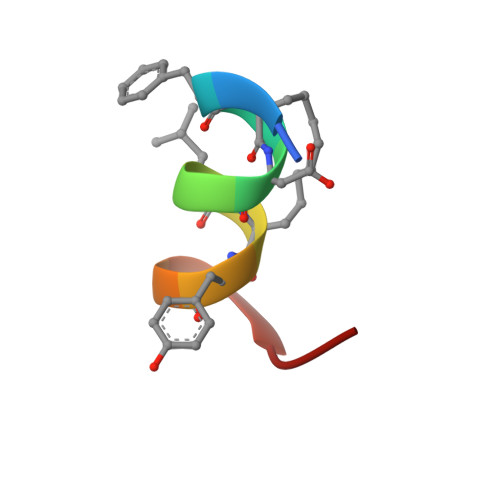
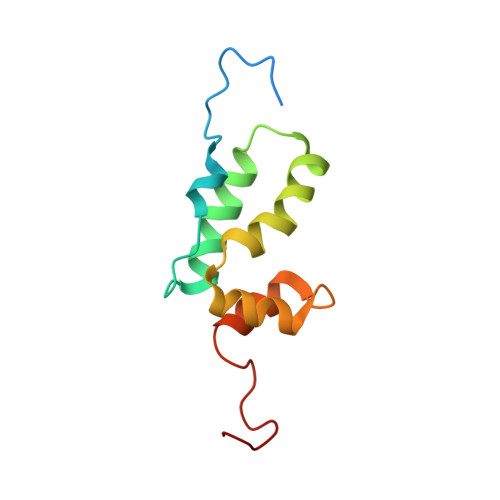
Solution structure of a hydrocarbon stapled peptide inhibitor in complex with monomeric C-terminal domain of HIV-1 capsid.
Bhattacharya, S., Zhang, H., Debnath, A.K., Cowburn, D.(2008) J Biological Chem 283: 16274-16278
- PubMed: 18417468 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.C800048200
- Primary Citation Related Structures:
2L6E - PubMed Abstract:
The human immunodeficiency virus type 1 (HIV-1) capsid protein plays a critical role in virus core particle assembly and is an important target for novel therapeutic strategies. In a previous study, we characterized the binding affinity of a hydrocarbon stapled helical peptide, NYAD-1, for the capsid protein (K(d) approximately 1 mum) and demonstrated its ability to penetrate the cell membrane (Zhang, H., Zhao, Q., Bhattacharya, S., Waheed, A. A., Tong, X., Hong, A., Heck, S., Goger, M., Cowburn, D., Freed, E. O., and Debnath, A. K. (2008) J. Mol. Biol. 378, 565-580). In cell-based assays, NYAD-1 colocalized with the Gag polyprotein during traffic to the plasma membrane and disrupted the formation of mature and immature virus particles in vitro systems. Here, we complement the cellular and biochemical data with structural characterization of the interactions between the capsid and a soluble peptide analogue, NYAD-13. Solution NMR methods were used to determine a high resolution structure of the complex between the inhibitor and a monomeric form of the C-terminal domain of the capsid protein (mCA-CTD). The intermolecular interactions are mediated by the packing of hydrophobic side chains at the buried interface and unperturbed by the presence of the olefinic chain on the solvent-exposed surface of the peptide. The results of the structural analysis provide valuable insight into the determinants for high affinity and selective inhibitors for HIV-1 particle assembly.
- New York Structural Biology Center, New York, New York 10027, USA.
Organizational Affiliation: