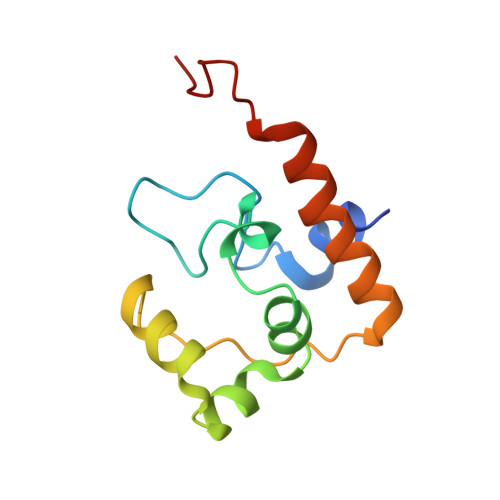
Sco proteins are involved in electron transfer processes
Banci, L., Bertini, I., Ciofi-Baffoni, S., Kozyreva, T., Mori, M., Wang, S.(2011) J Biol Inorg Chem 16: 391-403
- PubMed: 21181421 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-010-0735-x
- Primary Citation Related Structures:
2L4D - PubMed Abstract:
Sco proteins are widespread in eukaryotic and in many prokaryotic organisms. They have a thioredoxin-like fold and bind a single copper(I) or copper(II) ion through a CXXXC motif and a conserved His ligand, with both tight and weak affinities. They have been implicated in the assembly of the Cu(A) site of cytochrome c oxidase as copper chaperones and/or thioredoxins. In this work we have structurally characterized a Sco domain which is naturally fused with a typical electron transfer molecule, i.e., cytochrome c, in Pseudomonas putida. The thioredoxin-like Sco domain does not bind copper(II), binds copper(I) with weak affinity without involving the conserved His, and has redox properties consisting of a thioredoxin activity and of the ability of reducing copper(II) to copper(I), and iron(III) to iron(II) of the cytochrome c domain. These findings indicate that the His ligand coordination is the discriminating factor for introducing a metallochaperone function in a thioredoxin-like fold, typically responsible for electron transfer processes. A comparative structural analysis of the Sco domain from P. putida versus eukaryotic Sco proteins revealed structural determinants affecting the formation of a tight-affinity versus a weak-affinity copper binding site in Sco proteins.
- Magnetic Resonance Center CERM, University of Florence, Via Luigi Sacconi 6, 50019, Sesto Fiorentino, Florence, Italy.
Organizational Affiliation: