Structural and Functional Analysis of Transmembrane Segment VI of the NHE1 Isoform of the Na+/H+ Exchanger.
Tzeng, J., Lee, B.L., Sykes, B.D., Fliegel, L.(2010) J Biological Chem 285: 36656-36665
- PubMed: 20843797 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.161471
- Primary Citation Related Structures:
2L0E - PubMed Abstract:
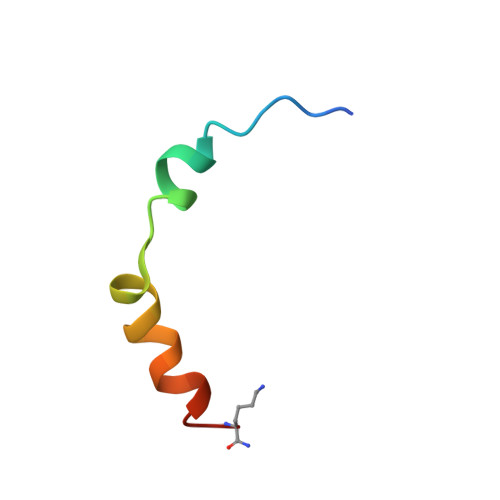
The Na(+)/H(+) exchanger isoform 1 is a ubiquitously expressed integral membrane protein. It resides on the plasma membrane of cells and regulates intracellular pH in mammals by extruding an intracellular H(+) in exchange for one extracellular Na(+). We characterized structural and functional aspects of the transmembrane segment (TM) VI (residues 227-249) by using cysteine scanning mutagenesis and high resolution NMR. Each residue of TM VI was mutated to cysteine in the background of the cysteineless NHE1 protein, and the sensitivity to water-soluble sulfhydryl-reactive compounds (2-(trimethylammonium)ethyl)methanethiosulfonate (MTSET) and (2-sulfonatoethyl)methanethiosulfonate (MTSES) was determined for those residues with significant activity remaining. Three residues were essentially inactive when mutated to Cys: Asp(238), Pro(239), and Glu(247). Of the remaining residues, proteins with the mutations N227C, I233C, and L243C were strongly inhibited by MTSET, whereas amino acids Phe(230), Gly(231), Ala(236), Val(237), Ala(244), Val(245), and Glu(248) were partially inhibited by MTSET. MTSES did not affect the activity of the mutant NHE1 proteins. The structure of a peptide representing TM VI was determined using high resolution NMR spectroscopy in dodecylphosphocholine micelles. TM VI contains two helical regions oriented at an approximate right angle to each other (residues 229-236 and 239-250) surrounding a central unwound region. This structure bears a resemblance to TM IV of the Escherichia coli protein NhaA. The results demonstrate that TM VI of NHE1 is a discontinuous pore-lining helix with residues Asn(227), Ile(233), and Leu(243) lining the translocation pore.
- Department of Biochemistry, University of Alberta, Edmonton, Alberta T6G 2H7, Canada.
Organizational Affiliation: