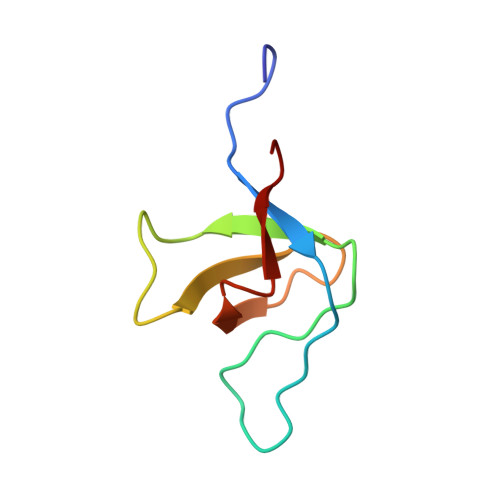
Recognition of tandem PxxP motifs as a unique Src homology 3-binding mode triggers pathogen-driven actin assembly
Aitio, O., Hellman, M., Kazlauskas, A., Vingadassalom, D.F., Leong, J.M., Saksela, K., Permi, P.(2010) Proc Natl Acad Sci U S A 107: 21743-21748
- PubMed: 21098279 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1010243107
- Primary Citation Related Structures:
2KXC - PubMed Abstract:
Src homology 3 (SH3) domains are globular protein interaction modules that regulate cell behavior. The classic SH3 ligand-binding site accommodates a hydrophobic PxxP motif and a positively charged specificity-determining residue. We have determined the NMR structure of insulin receptor tyrosine kinase substrate (IRTKS) SH3 domain in complex with a repeat from Escherichia coli-secreted protein F-like protein encoded on prophage U (EspF(U)), a translocated effector of enterohemorrhagic E. coli that commandeers the mammalian actin assembly machinery. EspF(U)-IRTKS interaction is among the highest affinity natural SH3 ligands. Our complex structure reveals a unique type of SH3 interaction based on recognition of tandem PxxP motifs in the ligand. Strikingly, the specificity pocket of IRTKS SH3 has evolved to accommodate a polyproline type II helical peptide analogously to docking of the canonical PxxP by the conserved IRTKS SH3 proline-binding pockets. This cooperative binding explains the high-affinity SH3 interaction and is required for EspF(U)-IRTKS interaction in mammalian cells as well as the formation of localized actin "pedestals" beneath bound bacteria. Importantly, tandem PxxP motifs are also found in mammalian ligands and have been shown to contribute to IRTKS SH3 recognition similarly.
- Program in Structural Biology and Biophysics, Institute of Biotechnology, University of Helsinki, FI-00014 Helsinki, Finland.
Organizational Affiliation: